Introduction to Difference-in-Differences (DiD) in Python
1. Overview
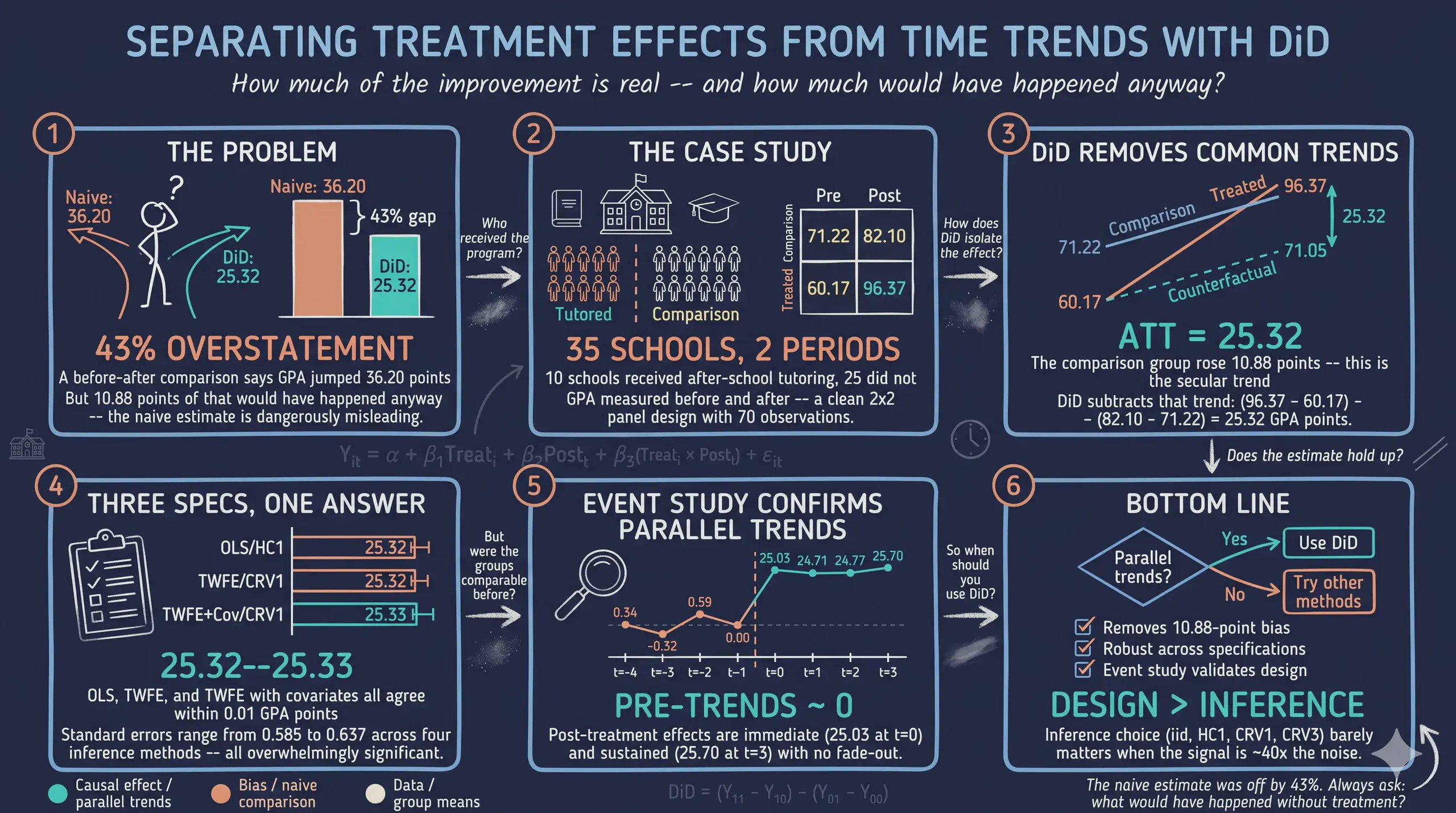
How much does an after-school tutoring program improve student performance? A school district implemented a new after-school tutoring program in 10 of its 35 high schools. After one year, the average GPA in tutored schools jumped from 60.17 to 96.37 — a staggering 36.20-point increase. Case closed?
Not quite. Over the same period, GPA also rose in the 25 schools that did not receive the program — from 71.22 to 82.10. Some of the improvement in tutored schools simply reflects a region-wide upward trend. Difference-in-Differences (DiD) strips away that common trend and reveals the tutoring program’s true causal effect: an ATT of approximately 25.32 GPA points.
This tutorial walks through DiD estimation in Python using PyFixest — a fast, Stata-flavored econometrics package — alongside Great Tables for publication-quality output. We use the simulated case study from Corral and Yang (2024), the same dataset used in the Stata companion tutorial.
1.1 Learning objectives
By the end of this tutorial, you will be able to:
- Explain why naive before-after comparisons overstate treatment effects
- Compute 2×2 DiD manually and via PyFixest’s
feols()function - Estimate DiD using multiple equivalent approaches with a unified formula syntax
- Compare inference under iid, HC1, CRV1, and CRV3 standard errors
- Build publication-quality regression tables with
etable()and Great Tables - Estimate and plot event study models with
i()for dynamic treatment effects
1.2 Study design
graph LR
subgraph "Case Study Setting"
A["<b>35 High Schools</b><br/>in One Region"]
B["<b>10 Treated Schools</b><br/>(tutoring program)"]
C["<b>25 Comparison Schools</b><br/>(no program)"]
A --> B
A --> C
end
subgraph "DiD Design"
D["<b>Pre-Program</b><br/>GPA at baseline"]
E["<b>Post-Program</b><br/>GPA after intervention"]
F["<b>DiD Estimate</b><br/>ATT = 25.32"]
D --> E --> F
end
subgraph "Estimation Methods"
G["<b>Manual 2x2</b><br/>Subtraction"]
H["<b>TWFE Regression</b><br/>PyFixest feols()"]
I["<b>Event Study</b><br/>Dynamic effects"]
G --> H --> I
end
C --> D
F --> G
style A fill:#6a9bcc,stroke:#141413,color:#fff
style B fill:#d97757,stroke:#141413,color:#fff
style C fill:#6a9bcc,stroke:#141413,color:#fff
style F fill:#00d4c8,stroke:#141413,color:#fff
style I fill:#00d4c8,stroke:#141413,color:#fff
The data has a clean panel structure: each of the 35 schools is observed in two time periods (pre and post), giving us 70 observations for the 2×2 design. A second dataset extends this to 8 periods (280 observations) for the event study analysis.
2. Setup and Imports
Install the required packages:
pip install pyfixest great_tables pandas matplotlib
Import the libraries:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pyfixest as pf
from great_tables import GT, md, style, loc
| Package | Purpose |
|---|---|
pyfixest |
Fast fixed-effects estimation with Stata-like formula syntax |
great_tables |
Publication-quality HTML/PNG tables from DataFrames |
pandas |
Data loading, manipulation, and summary statistics |
matplotlib |
Custom figure generation with dark theme styling |
Dark theme figure styling (click to expand)
# Site color palette
STEEL_BLUE = "#6a9bcc"
WARM_ORANGE = "#d97757"
NEAR_BLACK = "#141413"
TEAL = "#00d4c8"
# Dark theme palette
DARK_NAVY = "#0f1729"
GRID_LINE = "#1f2b5e"
LIGHT_TEXT = "#c8d0e0"
WHITE_TEXT = "#e8ecf2"
plt.rcParams.update({
"figure.facecolor": DARK_NAVY,
"axes.facecolor": DARK_NAVY,
"axes.edgecolor": DARK_NAVY,
"axes.linewidth": 0,
"axes.labelcolor": LIGHT_TEXT,
"axes.titlecolor": WHITE_TEXT,
"axes.spines.top": False,
"axes.spines.right": False,
"axes.spines.left": False,
"axes.spines.bottom": False,
"axes.grid": True,
"grid.color": GRID_LINE,
"grid.linewidth": 0.6,
"grid.alpha": 0.8,
"xtick.color": LIGHT_TEXT,
"ytick.color": LIGHT_TEXT,
"text.color": WHITE_TEXT,
"font.size": 12,
"legend.frameon": False,
"savefig.facecolor": DARK_NAVY,
})
3. Data Loading and Exploration
We load the 2×2 dataset directly from GitHub. This Stata .dta file contains 35 schools observed across 2 time periods:
url_did = "https://github.com/quarcs-lab/data-open/raw/master/isds/tutoring_did.dta"
df = pd.read_stata(url_did).astype(float)
print(df.shape)
print(df.dtypes)
(70, 7)
id float64
time float64
treated float64
post float64
txp float64
gpa float64
female_share float64
dtype: object
The dataset has 70 observations (35 schools × 2 periods) and 7 variables:
id— School identifier (1–35)time— Time period (1 = pre, 2 = post)treated— Treatment indicator (1 = received tutoring program)post— Post-period indicator (1 = after program implementation)txp— Interaction term (treated × post)gpa— Outcome: average GPA of low-income students (0–100 scale)female_share— Share of female students (covariate)
print(df.describe().round(2))
id time treated post txp gpa female_share
count 70.00 70.0 70.00 70.0 70.00 70.00 70.00
mean 18.00 1.5 0.29 0.5 0.14 77.12 0.53
std 10.17 0.5 0.46 0.5 0.35 10.88 0.03
min 1.00 1.0 0.00 0.0 0.00 59.39 0.47
25% 9.25 1.0 0.00 0.0 0.00 70.68 0.51
50% 18.00 1.5 0.00 0.5 0.00 76.27 0.53
75% 26.75 2.0 1.00 1.0 0.00 82.66 0.55
max 35.00 2.0 1.00 1.0 1.00 99.15 0.57
A crosstab confirms the balanced 2×2 design:
ct = pd.crosstab(df["treated"], df["post"], margins=True)
print(ct)
Pre (0) Post (1) Total
Comparison (0) 25 25 50
Treated (1) 10 10 20
Total 35 35 70
We have 10 treated schools observed in 2 periods (20 observations) and 25 comparison schools (50 observations). This is a perfectly balanced panel — every school appears exactly once in each period.
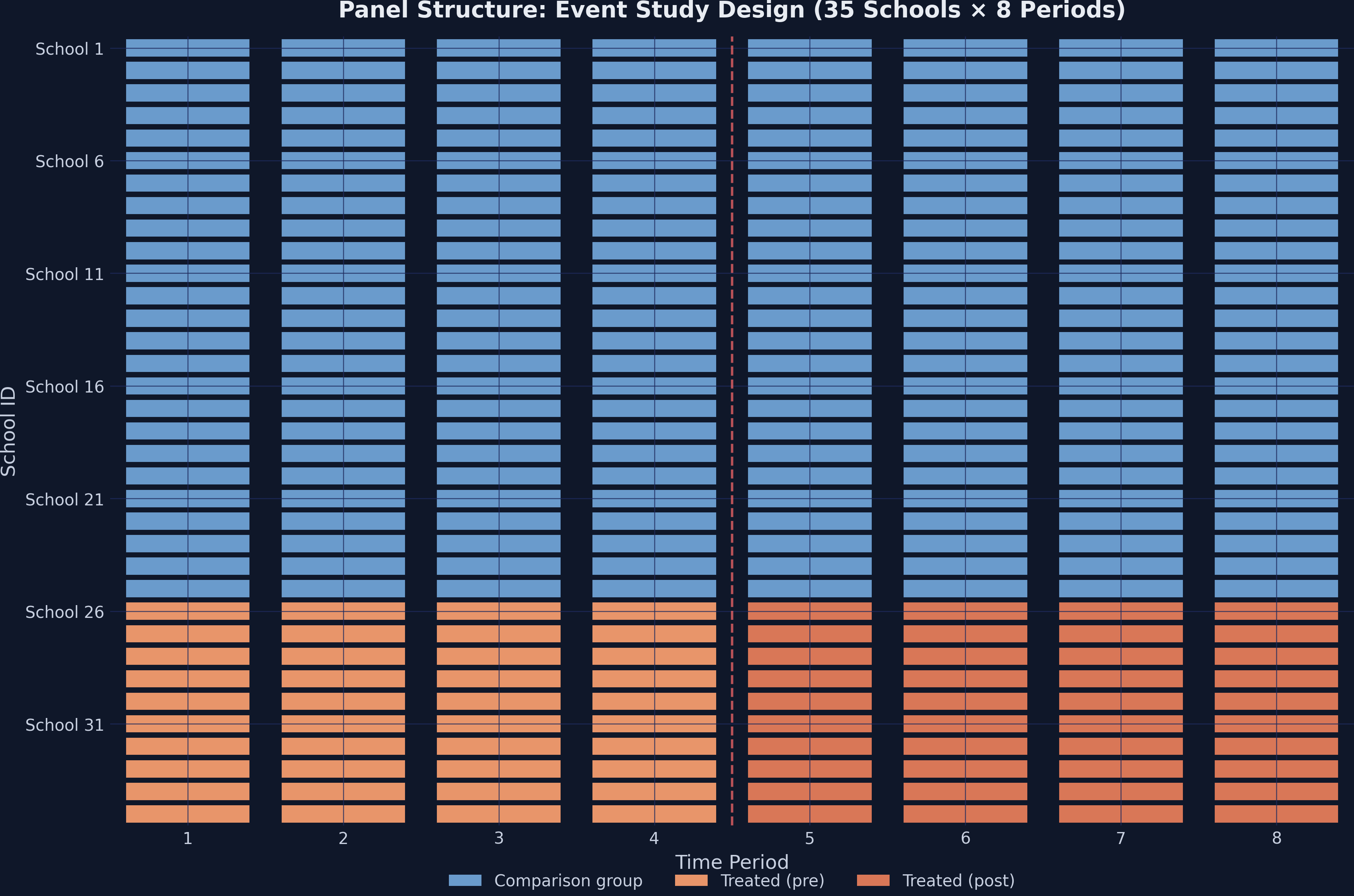
3.1 Panel structure visualization
The heatmap below shows the treatment assignment across schools and time. Steel blue cells represent the comparison group, while orange cells indicate treated schools in the post-program period.
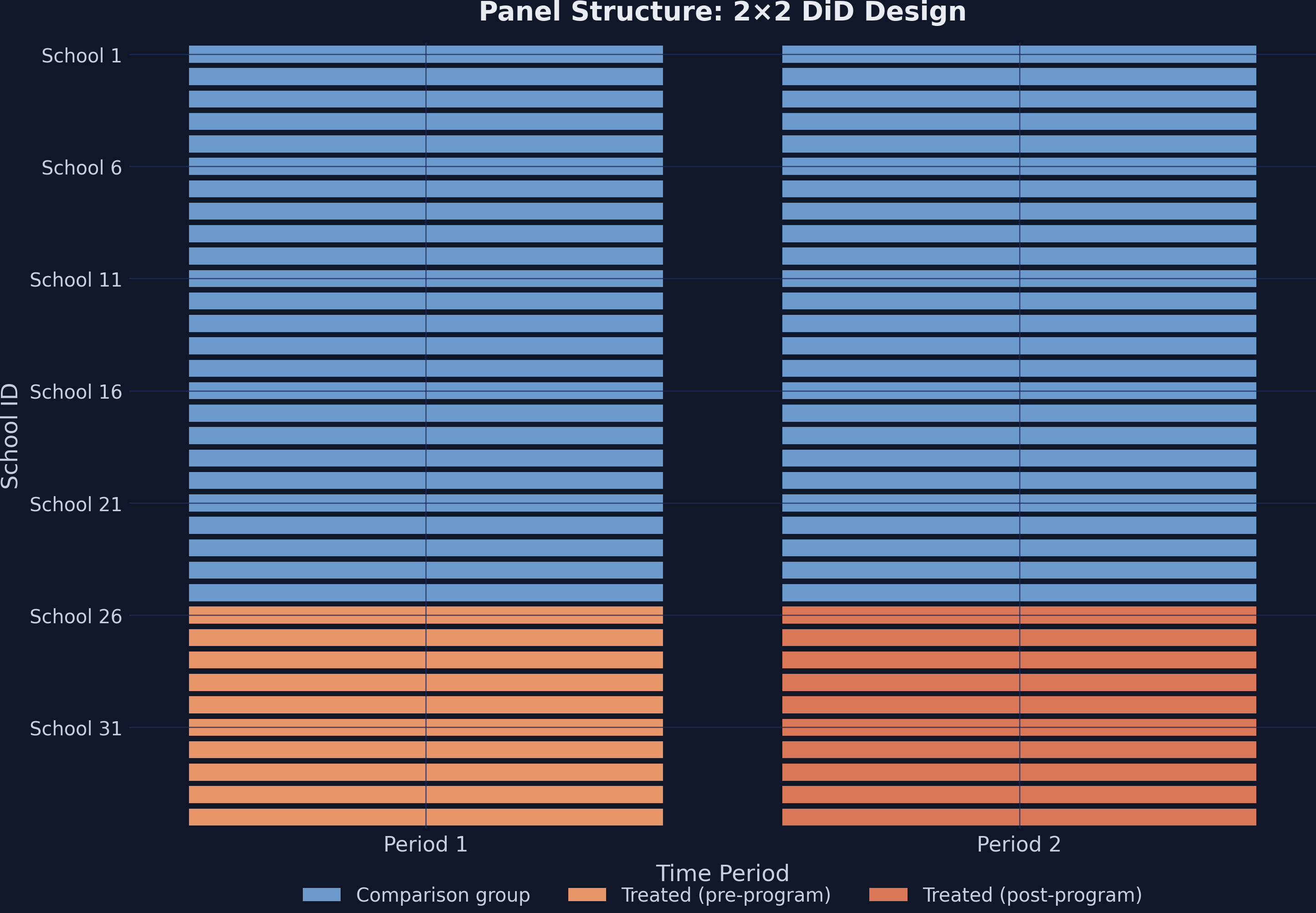
This is a clean 2×2 design: treatment timing is simultaneous (all 10 schools receive the program at the same time), and no school switches treatment status.
4. The Problem with Naive Comparisons
The most intuitive approach to measuring the program’s effect is a simple before-after comparison for the treated schools:
treated_means = df[df["treated"] == 1].groupby("post")["gpa"].mean()
print(f"Pre-program: {treated_means[0]:.2f}")
print(f"Post-program: {treated_means[1]:.2f}")
print(f"Naive change: {treated_means[1] - treated_means[0]:.2f}")
Pre-program: 60.17
Post-program: 96.37
Naive change: 36.20
The naive estimate says the program boosted GPA by 36.20 points. But this ignores everything else that may have changed over the same period — curriculum reforms, new textbooks, regional economic shifts, or simply students maturing. Any of these factors could drive GPA upward in all schools, not just the treated ones.
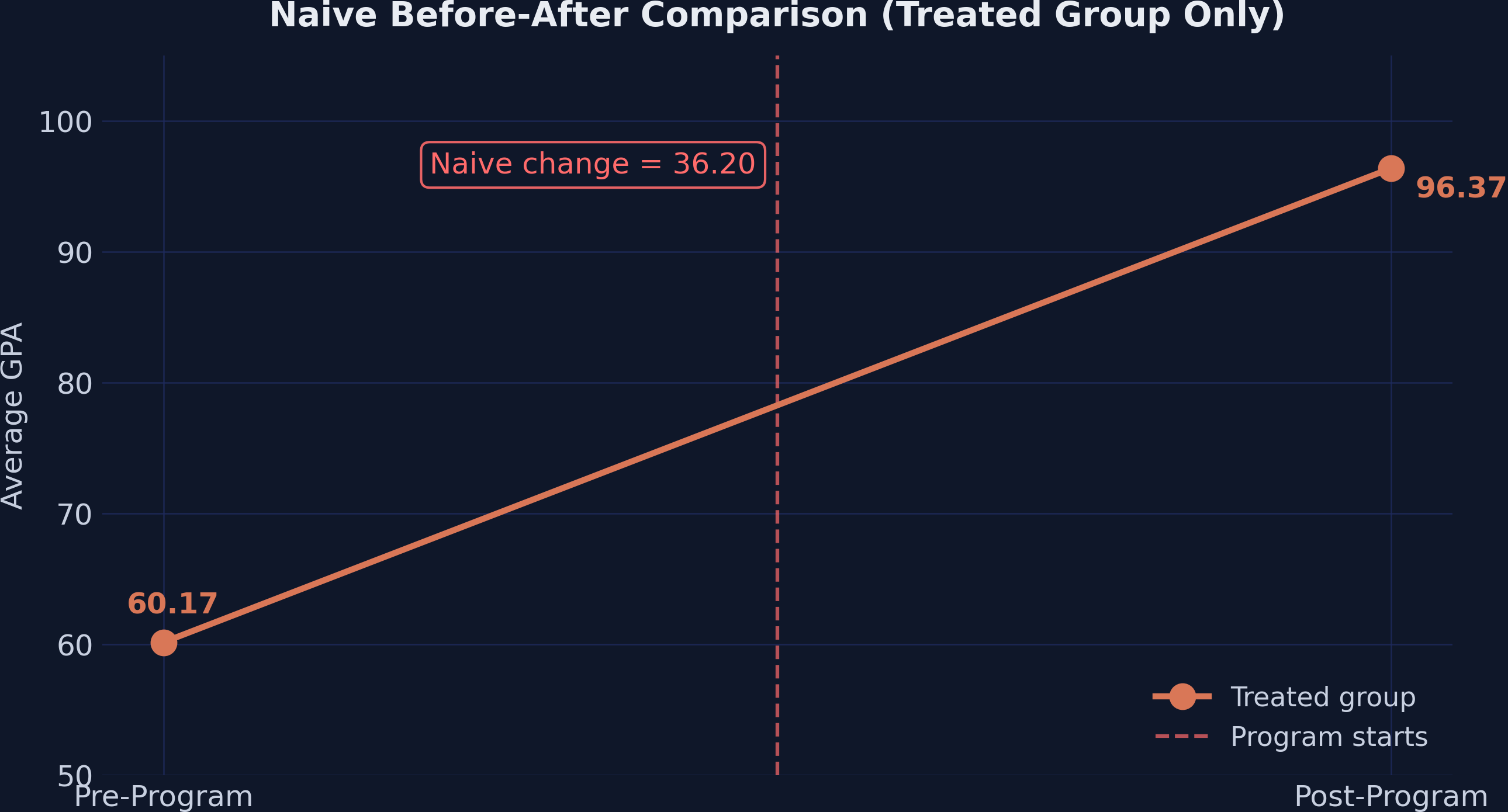
The naive approach overstates the effect by conflating the treatment effect with time trends that would have occurred regardless of the program.
5. The DiD Design: Using a Comparison Group
The key insight of DiD is to use the comparison group as a mirror for what would have happened to the treated schools without the program. We compute all four group means:
means = df.groupby(["treated", "post"])["gpa"].mean()
pre_control = means[(0, 0)] # 71.22
post_control = means[(0, 1)] # 82.10
pre_treated = means[(1, 0)] # 60.17
post_treated = means[(1, 1)] # 96.37
Group means:
Comparison Pre: 71.22
Comparison Post: 82.10
Treated Pre: 60.17
Treated Post: 96.37
The comparison schools' GPA rose by 10.88 points (from 71.22 to 82.10) — this is the secular trend. We assume the treated schools would have experienced the same trend absent the program. This gives us the counterfactual:
counterfactual = pre_treated + (post_control - pre_control)
did_estimate = post_treated - counterfactual
print(f"Counterfactual: {pre_treated:.2f} + ({post_control:.2f} - {pre_control:.2f}) = {counterfactual:.2f}")
print(f"DiD estimate: {post_treated:.2f} - {counterfactual:.2f} = {did_estimate:.2f}")
Counterfactual: 60.17 + (82.10 - 71.22) = 71.05
DiD estimate: 96.37 - 71.05 = 25.32
The causal effect of the tutoring program is 25.32 GPA points — not 36.20. The naive approach overstated the effect by 43% because it attributed the 10.88-point common trend entirely to the program.
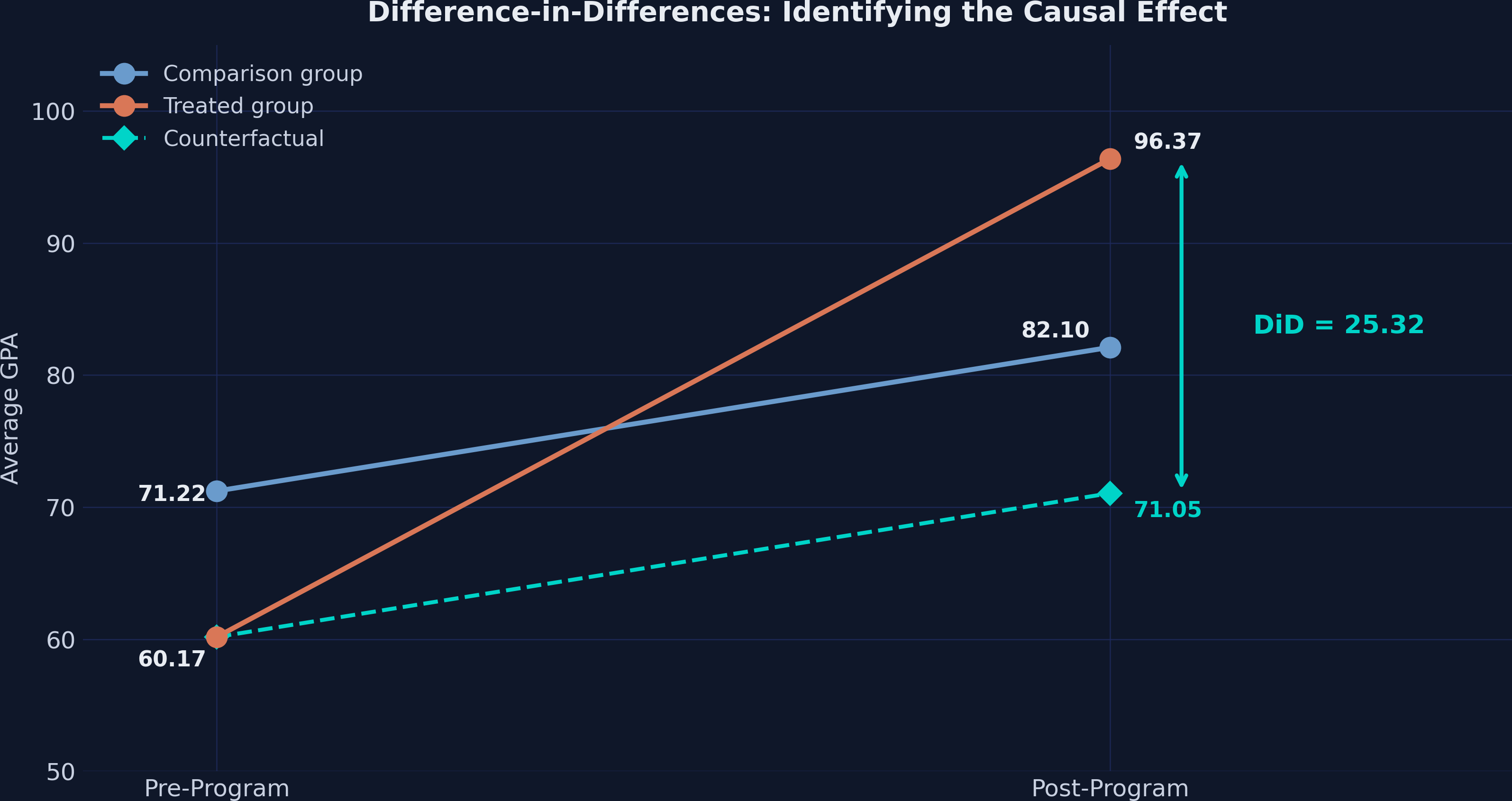
5.1 The parallel trends assumption
DiD rests on one critical assumption: parallel trends. In the absence of treatment, treated and comparison groups would have followed the same trajectory over time. Formally:
$$E[Y_{i,1}(0) - Y_{i,0}(0) \mid D=1] = E[Y_{i,1}(0) - Y_{i,0}(0) \mid D=0]$$
In words: the change in potential untreated outcomes is the same for both groups. Think of it like two runners on parallel tracks — they may start at different positions (treated schools have lower baseline GPA), but they run at the same pace. If one runner suddenly speeds up after receiving coaching, the difference between their new speed and the other runner’s speed measures the coaching effect.
Note what parallel trends does not require: the two groups do not need the same level of GPA, only the same trend. This is why DiD is powerful — it naturally handles time-invariant differences between groups (like school quality or student demographics).
5.2 SUTVA
The Stable Unit Treatment Value Assumption (SUTVA) requires that one school’s treatment does not affect another school’s outcome. If untreated schools lost students to tutored schools, or if tutored schools drew resources away from comparison schools, the DiD estimate would be biased. In this setting, schools serve distinct geographic catchments, making spillovers unlikely.
6. Manual DiD Calculation
We can organize the four group means into a 2×2 table and compute the DiD as a double difference:
means_table = df.groupby(["treated", "post"])["gpa"].mean().unstack()
means_table["Difference"] = means_table[1.0] - means_table[0.0]
print(means_table.round(2))
Pre (0) Post (1) Difference
Comparison (0) 71.22 82.10 10.88
Treated (1) 60.17 96.37 36.20
The DiD formula takes the difference of differences:
$$DiD = \Big(E[Y_{i,1} \mid D=1] - E[Y_{i,0} \mid D=1]\Big) - \Big(E[Y_{i,1} \mid D=0] - E[Y_{i,0} \mid D=0]\Big)$$
Plugging in the numbers:
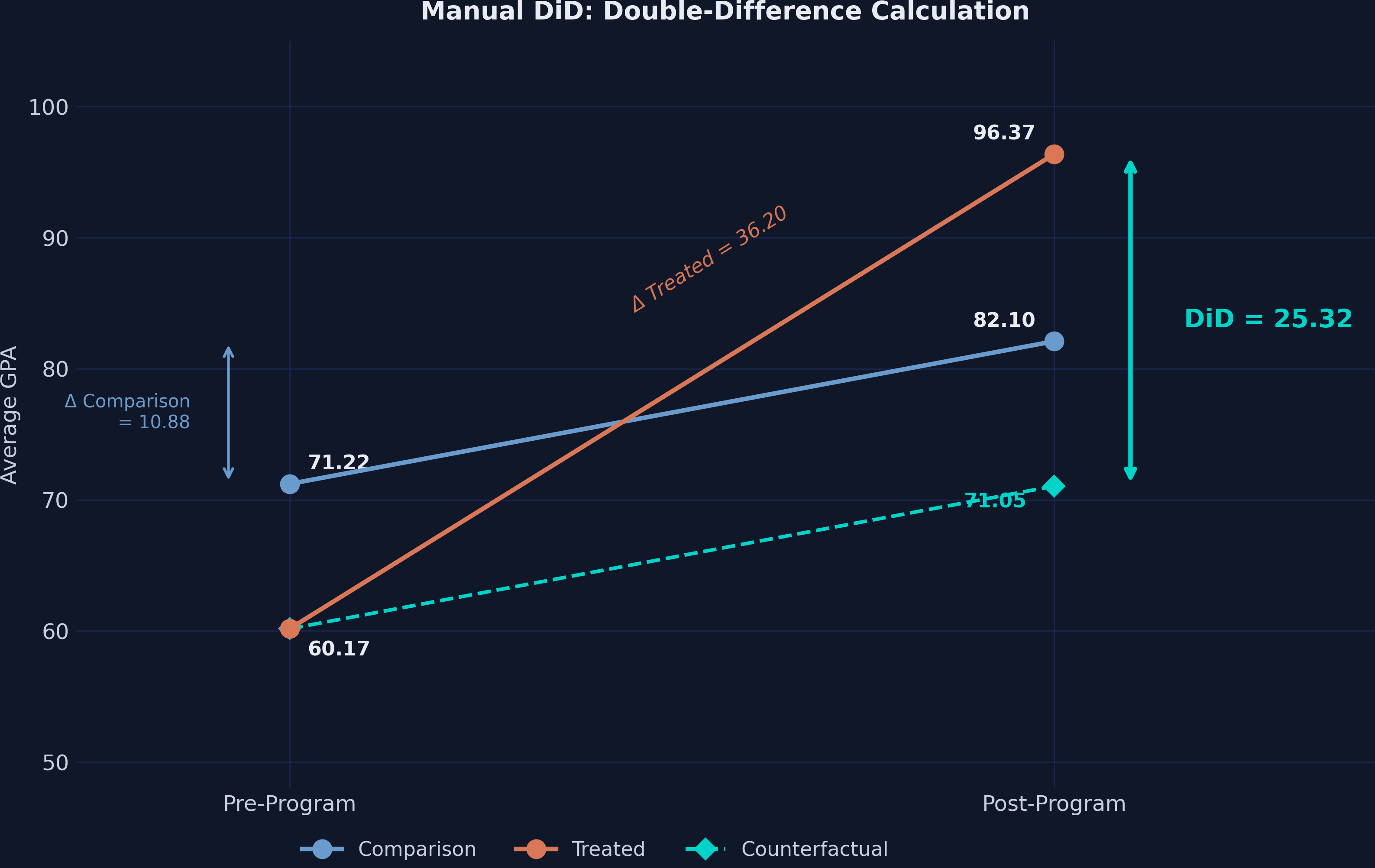
$$DiD = (96.37 - 60.17) - (82.10 - 71.22) = 36.20 - 10.88 = 25.32$$
Think of it this way: the treated schools improved by 36.20 points, but 10.88 of those points would have happened anyway (as evidenced by the comparison group). The remaining 25.32 points is the causal effect of the tutoring program.
Going back to the runner analogy: the treated runner sped up by 36.20 units while the comparison runner sped up by 10.88. The coaching effect is the extra 25.32 units of speed that only the coached runner gained.
7. DiD via Regression
7.1 Classical OLS with interaction
The manual calculation is equivalent to an OLS regression with the treatment indicator, time indicator, and their interaction:
$$Y_{it} = \alpha + \beta_1 \text{Treat}_i + \beta_2 \text{Post}_t + \beta_3 (\text{Treat}_i \times \text{Post}_t) + \varepsilon_{it}$$
Where:
- $\alpha$ is the comparison group’s pre-period mean (intercept)
- $\beta_1$ captures the baseline difference between groups
- $\beta_2$ captures the common time trend
- $\beta_3$ is the DiD estimate — the causal effect of treatment
In PyFixest, the feols() function handles this with a familiar formula syntax:
fit_ols = pf.feols("gpa ~ treated + post + txp", data=df, vcov="HC1")
print(fit_ols.summary())
Estimation: OLS
Dep. var.: gpa, Fixed effects: 0
Inference: HC1
Observations: 70
| Coefficient | Estimate | Std. Error | t value | Pr(>|t|) | 2.5% | 97.5% |
|:--------------|-----------:|-------------:|----------:|-----------:|--------:|--------:|
| Intercept | 71.215 | 0.218 | 326.123 | 0.000 | 70.779 | 71.651 |
| treated | -11.049 | 0.288 | -38.388 | 0.000 | -11.624 | -10.475 |
| post | 10.886 | 0.339 | 32.116 | 0.000 | 10.209 | 11.563 |
| txp | 25.315 | 0.615 | 41.164 | 0.000 | 24.087 | 26.543 |
---
RMSE: 1.15 R2: 0.989
Every coefficient maps directly to our group means:
- Intercept (71.22) — Comparison group pre-period mean
- treated (−11.05) — Treated schools start 11 points below comparison schools
- post (10.89) — Common time trend (comparison group’s improvement)
- txp (25.32) — The DiD estimate, matching our manual calculation
The vcov="HC1" option requests heteroskedasticity-robust (White) standard errors, the most common choice for cross-sectional data.
7.2 TWFE with fixed effects
A more flexible approach absorbs school-level and time-level heterogeneity using two-way fixed effects (TWFE). PyFixest uses the | pipe syntax to specify absorbed fixed effects:
$$Y_{it} = \beta_3 (\text{Treat}_i \times \text{Post}_t) + \gamma_i + \vartheta_t + \varepsilon_{it}$$
Here $\gamma_i$ are school fixed effects (absorbing all time-invariant school characteristics) and $\vartheta_t$ are time fixed effects (absorbing all common time shocks). Since treated is perfectly collinear with $\gamma_i$ and post is perfectly collinear with $\vartheta_t$, only the interaction term txp remains:
fit_twfe = pf.feols("gpa ~ txp | id + time", data=df, vcov={"CRV1": "id"})
print(fit_twfe.summary())
Estimation: OLS
Dep. var.: gpa, Fixed effects: id+time
Inference: CRV1
Observations: 70
| Coefficient | Estimate | Std. Error | t value | Pr(>|t|) | 2.5% | 97.5% |
|:--------------|-----------:|-------------:|----------:|-----------:|-------:|--------:|
| txp | 25.315 | 0.585 | 43.265 | 0.000 | 24.126 | 26.504 |
---
RMSE: 0.788 R2: 0.995 R2 Within: 0.981
The estimate is unchanged: 25.315. But the standard errors now use CRV1 (cluster-robust variance) clustered at the school level — the appropriate choice when treatment varies at the school level and observations within the same school are correlated.
The formula "gpa ~ txp | id + time" is one of PyFixest’s key strengths: everything to the left of | is estimated, everything to the right is absorbed. No need to manually create dummy variables.
7.3 TWFE with covariate
We can add female_share as a time-varying covariate to check robustness:
fit_cov = pf.feols("gpa ~ txp + female_share | id + time", data=df,
vcov={"CRV1": "id"})
print(fit_cov.summary())
Estimation: OLS
Dep. var.: gpa, Fixed effects: id+time
Inference: CRV1
Observations: 70
| Coefficient | Estimate | Std. Error | t value | Pr(>|t|) | 2.5% | 97.5% |
|:--------------|-----------:|-------------:|----------:|-----------:|--------:|--------:|
| txp | 25.328 | 0.605 | 41.881 | 0.000 | 24.099 | 26.557 |
| female_share | -3.216 | 8.700 | -0.370 | 0.714 | -20.898 | 14.465 |
---
RMSE: 0.785 R2: 0.995 R2 Within: 0.982
Adding female_share barely changes the DiD estimate (25.315 → 25.328, a shift of just 0.013). The covariate itself is statistically insignificant (p = 0.714), confirming that the two-way fixed effects already capture the relevant variation. This is reassuring — the treatment effect estimate is robust to the inclusion of observable covariates.
7.4 Programmatic access to results
PyFixest provides tidy methods for extracting specific quantities — useful for post-estimation workflows and building custom tables:
print(f"Coefficient: {fit_twfe.coef().values[0]:.4f}")
print(f"Std. Error: {fit_twfe.se().values[0]:.4f}")
print(f"t-statistic: {fit_twfe.tstat().values[0]:.4f}")
print(f"p-value: {fit_twfe.pvalue().values[0]:.4f}")
print(f"95% CI: [{fit_twfe.confint().values[0, 0]:.2f}, {fit_twfe.confint().values[0, 1]:.2f}]")
Coefficient: 25.3149
Std. Error: 0.5851
t-statistic: 43.2655
p-value: 0.0000
95% CI: [24.13, 26.50]
The .tidy() method returns a full DataFrame of results:
print(fit_twfe.tidy())
Estimate Std. Error t value Pr(>|t|) 2.5% 97.5%
Coefficient
txp 25.314897 0.585106 43.265472 0.0 24.125818 26.503976
7.5 Comparison across specifications
All three specifications produce essentially the same DiD estimate:
| Method | Estimate | Std. Error | 95% CI | SE Type |
|---|---|---|---|---|
| OLS / HC1 | 25.315 | 0.615 | [24.09, 26.54] | Heteroskedasticity-robust |
| TWFE / CRV1 | 25.315 | 0.585 | [24.13, 26.50] | Cluster-robust (school) |
| TWFE + Cov / CRV1 | 25.328 | 0.605 | [24.10, 26.56] | Cluster-robust (school) |
The point estimates range from 25.315 to 25.328 — a difference of just 0.013 GPA points. The design (treatment assignment, fixed effects) does the heavy lifting; the choice of specification has negligible impact on the estimate.
8. Inference Comparison
One of PyFixest’s strengths is the ability to quickly compare different inference approaches on the same model. Here we estimate the TWFE model four times, each with a different variance-covariance estimator:
vcov_types = {
"iid": "iid",
"HC1": "HC1",
"CRV1": {"CRV1": "id"},
"CRV3": {"CRV3": "id"},
}
for label, vcov_spec in vcov_types.items():
fit_tmp = pf.feols("gpa ~ txp | id + time", data=df, vcov=vcov_spec)
tidy = fit_tmp.tidy()
txp_row = tidy[tidy.index == "txp"].iloc[0]
print(f" {label:5s}: SE = {txp_row['Std. Error']:.4f}, "
f"t = {txp_row['t value']:.2f}, "
f"p = {txp_row['Pr(>|t|)']:.4f}")
iid : SE = 0.6071, t = 41.70, p = 0.0000
HC1 : SE = 0.5852, t = 43.26, p = 0.0000
CRV1 : SE = 0.5851, t = 43.27, p = 0.0000
CRV3 : SE = 0.6373, t = 39.72, p = 0.0000
| SE Type | Description | SE | t-stat |
|---|---|---|---|
| iid | Classical (assumes homoskedasticity) | 0.607 | 41.70 |
| HC1 | Heteroskedasticity-robust (White) | 0.585 | 43.26 |
| CRV1 | Cluster-robust at school level | 0.585 | 43.27 |
| CRV3 | Bias-corrected cluster-robust (Bell-McCaffrey) | 0.637 | 39.72 |
- iid assumes constant variance — a strong assumption rarely justified in practice
- HC1 allows for heteroskedasticity but not within-cluster correlation
- CRV1 is the workhorse for panel data: it accounts for arbitrary within-school correlation
- CRV3 applies a small-sample bias correction, producing the most conservative SEs
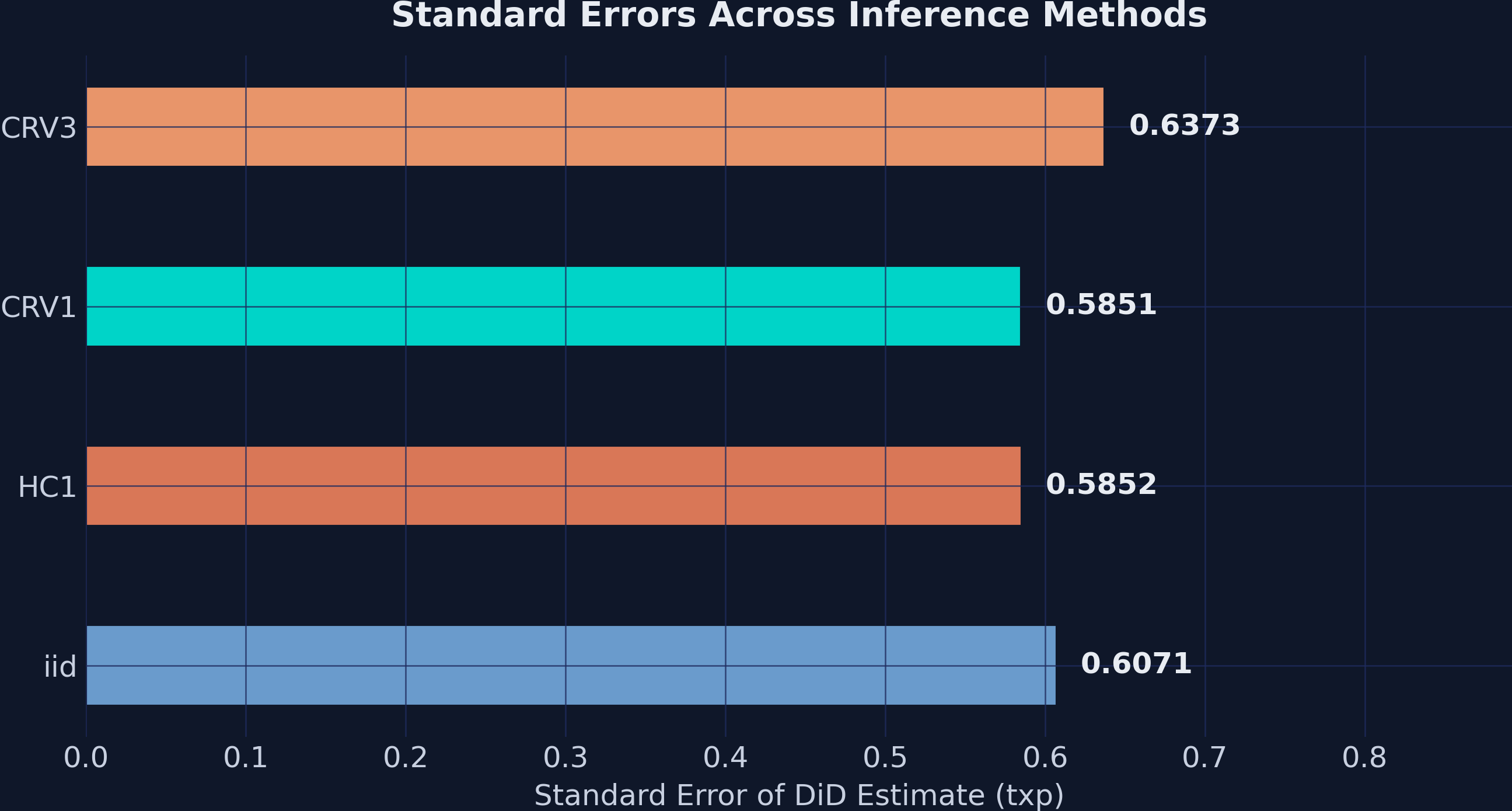
The key takeaway: inference choice matters less than research design. Standard errors range from 0.585 to 0.637, but all four methods produce p-values that are essentially zero. When the treatment effect is 25 GPA points and the largest SE is 0.64, the t-statistic is still above 39. The signal-to-noise ratio is so strong that the choice of variance estimator is practically irrelevant here.
In applications with smaller effects or fewer clusters, the choice between CRV1 and CRV3 can make the difference between statistical significance and not. With only 35 clusters, CRV3 is the safer default.
9. Publication-Quality Tables with etable() and Great Tables
9.1 Stepwise specifications with csw0()
PyFixest’s csw0() operator lets you estimate multiple specifications in a single call. The csw0 (“cumulative stepwise from zero”) starts with a baseline model and progressively adds covariates:
fit_multi = pf.feols("gpa ~ txp + csw0(female_share) | id + time",
data=df, vcov={"CRV1": "id"})
This single line estimates two models: (1) gpa ~ txp | id + time and (2) gpa ~ txp + female_share | id + time. In Stata, you would need two separate regression commands; PyFixest handles it in one formula.
9.2 etable() output
The etable() method generates a publication-style regression table as a Great Tables object:
print(fit_multi.etable())
(1) (2)
txp 25.315*** (0.585) 25.328*** (0.605)
female_share -3.216 (8.700)
---
FE: time x x
FE: id x x
Observations 70 70
S.E. type by: id by: id
R² 0.995 0.995
R² Within 0.981 0.982
The etable() output shows significance stars, standard errors in parentheses, fixed effects indicators, and model diagnostics — all formatted for immediate inclusion in a paper.
9.3 Custom Great Tables table
For full control over formatting, we can build a table from the .tidy() DataFrames:
rows = []
for name, fit in [("(1) OLS", fit_ols),
("(2) TWFE", fit_twfe),
("(3) TWFE + Cov", fit_cov)]:
tidy = fit.tidy()
txp_row = tidy[tidy.index == "txp"].iloc[0]
rows.append({
"Model": name,
"Estimate": txp_row["Estimate"],
"Std. Error": txp_row["Std. Error"],
"t value": txp_row["t value"],
"p-value": txp_row["Pr(>|t|)"],
"95% CI Lower": txp_row["2.5%"],
"95% CI Upper": txp_row["97.5%"],
"N": fit._N,
})
gt_df = pd.DataFrame(rows)
gt_table = (
GT(gt_df)
.tab_header(
title=md("**Table 2: DiD Estimates Across Specifications**"),
subtitle="Dependent variable: GPA"
)
.fmt_number(columns=["Estimate", "Std. Error", "t value",
"95% CI Lower", "95% CI Upper"], decimals=3)
.fmt_number(columns=["p-value"], decimals=4)
.fmt_integer(columns=["N"])
.tab_source_note(
"Notes: (1) OLS with HC1 robust SE. (2) TWFE with CRV1 "
"clustered at school level. (3) TWFE with female_share covariate and CRV1."
)
)
gt_table.save("did101_table2.png")
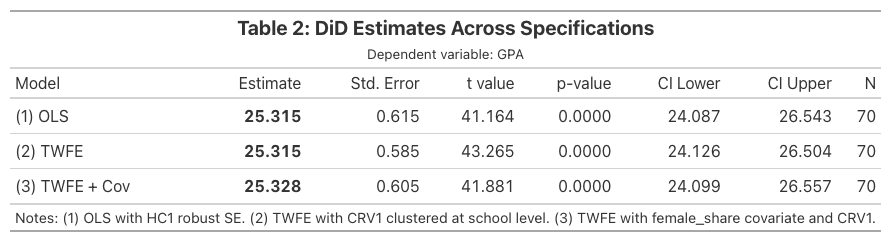
Great Tables provides fine-grained control over number formatting (.fmt_number()), column labels (.cols_label()), headers (.tab_header()), and styling (.tab_style()). The .save() method exports to PNG using a headless browser.
9.4 Exporting LaTeX tables for manuscripts
When submitting to academic journals, you need LaTeX-formatted tables rather than HTML or PNG. PyFixest’s etable() can generate publication-ready LaTeX directly by setting type="tex". The output uses booktabs for clean horizontal rules and threeparttable for properly aligned footnotes — the standard format expected by most economics and social science journals.
latex_output = pf.etable(
[fit_ols, fit_twfe, fit_cov],
type="tex",
labels={
"txp": "Treatment $\\times$ Post",
"treated": "Treatment",
"post": "Post",
"female_share": "Female Share",
"Intercept": "Constant",
},
notes="Standard errors in parentheses. * p<0.05, ** p<0.01, *** p<0.001.",
)
print(latex_output)
\begin{threeparttable}
\begin{tabular}{lcccc}
\toprule
& \multicolumn{3}{c}{gpa} \\
\cmidrule(lr){2-4}
& (1) & (2) & (3) \\
\midrule
Treatment & \makecell{-11.049*** \\ (0.288)} & & \\
Post & \makecell{10.886*** \\ (0.339)} & & \\
Treatment:Post & \makecell{25.315*** \\ (0.615)} & \makecell{25.315*** \\ (0.585)} & \makecell{25.328*** \\ (0.605)} \\
Female Share & & & \makecell{-3.216 \\ (8.700)} \\
Constant & \makecell{71.215*** \\ (0.218)} & & \\
\midrule
id & - & x & x \\
time & - & x & x \\
\midrule
Observations & 70 & 70 & 70 \\
S.E. type & hetero & by: id & by: id \\
$R^2$ & 0.989 & 0.995 & 0.995 \\
$R^2$ Within & - & 0.981 & 0.982 \\
\bottomrule
\end{tabular}
\footnotesize Standard errors in parentheses. * p<0.05, ** p<0.01, *** p<0.001.
\end{threeparttable}
To save the table directly to a .tex file that you can \input{} in your manuscript, use the file_name parameter:
pf.etable(
[fit_ols, fit_twfe, fit_cov],
type="tex",
labels={
"txp": "Treatment $\\times$ Post",
"treated": "Treatment",
"post": "Post",
"female_share": "Female Share",
"Intercept": "Constant",
},
notes="Standard errors in parentheses. * p<0.05, ** p<0.01, *** p<0.001.",
file_name="did101_table2.tex",
)
This saves the file to did101_table2.tex. In your LaTeX manuscript, include it with:
\begin{table}[htbp]
\centering
\caption{DiD Estimates Across Specifications}
\label{tab:did-results}
\input{did101_table2.tex}
\end{table}
The labels dictionary maps internal variable names to publication-friendly labels (e.g., "txp" becomes "Treatment $\times$ Post"). The notes parameter adds a footnote below the table. Your LaTeX document needs the booktabs, makecell, and threeparttable packages in the preamble:
\usepackage{booktabs}
\usepackage{makecell}
\usepackage{threeparttable}
10. Coefficient Comparison
A coefficient plot provides a visual comparison of the DiD estimate across specifications, including 95% confidence intervals:
fig, ax = plt.subplots(figsize=(9, 5))
model_names = ["(1) OLS\nHC1", "(2) TWFE\nCRV1", "(3) TWFE+Cov\nCRV1"]
estimates = [fit.tidy().loc["txp", "Estimate"] for fit in [fit_ols, fit_twfe, fit_cov]]
ci_lower = [fit.tidy().loc["txp", "2.5%"] for fit in [fit_ols, fit_twfe, fit_cov]]
ci_upper = [fit.tidy().loc["txp", "97.5%"] for fit in [fit_ols, fit_twfe, fit_cov]]
ax.errorbar(estimates, range(3), xerr=[[e-l for e,l in zip(estimates, ci_lower)],
[u-e for e,u in zip(estimates, ci_upper)]],
fmt="o", color=TEAL, markersize=10, capsize=6, elinewidth=2)
ax.set_xlabel("DiD Estimate (txp coefficient)")
ax.set_title("Coefficient Comparison Across Specifications")
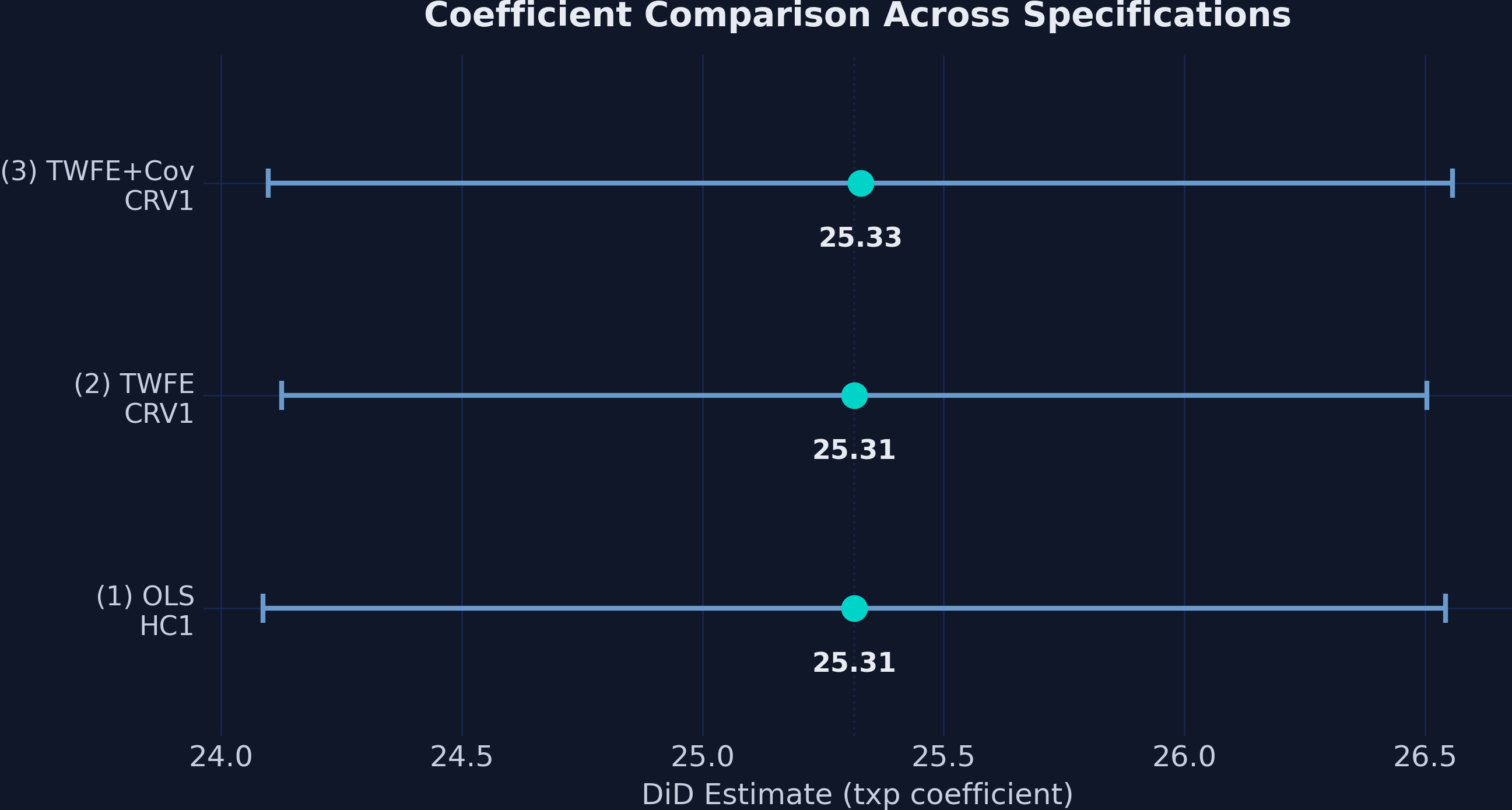
The near-identical point estimates and overlapping confidence intervals across all three specifications reinforce that the DiD estimate is robust. The point estimates span a range of just 0.013 GPA points (25.315 to 25.328), demonstrating remarkable stability regardless of whether we include school fixed effects, time fixed effects, or time-varying covariates.
11. Event Study: Dynamic Treatment Effects
11.1 Loading the event study data
The 2×2 design tells us whether the program had an effect, but not when the effect kicked in or whether it grew or faded over time. The event study dataset extends the analysis to 8 time periods (4 pre-treatment and 4 post-treatment):
url_event = "https://github.com/quarcs-lab/data-open/raw/master/isds/tutoring_didevent.dta"
df_event = pd.read_stata(url_event).astype(float)
print(f"Shape: {df_event.shape}")
Shape: (280, 8)
The new variable timeToTreat measures periods relative to treatment onset for treated schools: −4 through −1 are pre-treatment periods, 0 through 3 are post-treatment. Untreated schools have NaN for this variable (they are never treated).
print(df_event["timeToTreat"].value_counts().sort_index())
timeToTreat
-4.0 10
-3.0 10
-2.0 10
-1.0 10
0.0 10
1.0 10
2.0 10
3.0 10
Each of the 10 treated schools contributes one observation per relative time period, giving 10 observations at each event time.
11.2 The event study specification
The event study model replaces the single txp interaction with a full set of event-time indicators, one for each period relative to treatment. We omit one period (the reference period, $t = -1$) to avoid perfect collinearity:
$$Y_{it} = \alpha + \sum_{j=-m}^{q} \theta_j \cdot \text{treat}_{it}(t = k + j) + \gamma_i + \vartheta_t + \varepsilon_{it}$$
Each $\theta_j$ measures the treatment effect at event time $j$:
- Pre-treatment coefficients ($j < 0$): These should be near zero if parallel trends holds. Significant pre-treatment coefficients would indicate that treated schools were already diverging before the program — a red flag for the DiD design.
- Post-treatment coefficients ($j \geq 0$): These capture the dynamic treatment effect at each lag after the program starts.
11.3 Estimation with i()
PyFixest’s i() function creates factor (indicator) variables with a specified reference level — perfect for event study designs:
df_event["timeToTreat"] = df_event["timeToTreat"].fillna(-99)
fit_event = pf.feols("gpa ~ i(timeToTreat, ref=-1) | id + time",
data=df_event, vcov={"CRV1": "id"})
print(fit_event.summary())
Estimation: OLS
Dep. var.: gpa, Fixed effects: id+time
Inference: CRV1
Observations: 280
| Coefficient | Estimate | Std. Error | t value | Pr(>|t|) | 2.5% | 97.5% |
|:--------------|-----------:|-------------:|----------:|-----------:|-------:|--------:|
| t = -4 | 0.342 | 0.401 | 0.852 | 0.400 | -0.474 | 1.157 |
| t = -3 | -0.322 | 0.441 | -0.730 | 0.471 | -1.219 | 0.575 |
| t = -2 | 0.593 | 0.423 | 1.401 | 0.170 | -0.267 | 1.454 |
| t = 0 | 25.028 | 0.445 | 56.232 | 0.000 | 24.123 | 25.932 |
| t = 1 | 24.705 | 0.559 | 44.174 | 0.000 | 23.569 | 25.842 |
| t = 2 | 24.768 | 0.739 | 33.534 | 0.000 | 23.267 | 26.270 |
| t = 3 | 25.701 | 0.797 | 32.268 | 0.000 | 24.083 | 27.320 |
---
RMSE: 1.134 R2: 0.991 R2 Within: 0.961
Note: Coefficient names simplified for readability. PyFixest outputs C(timeToTreat, contr.treatment(base=-1))[T.X.0] notation, where X is the relative time period.
The i(timeToTreat, ref=-1) syntax tells PyFixest to create indicator variables for each unique value of timeToTreat, using $t = -1$ as the reference period (coefficient normalized to zero). We fill NaN values with −99 for untreated schools — this creates a dummy that gets absorbed by the school fixed effects, keeping the remaining coefficients interpretable.
11.4 Event study plot
The event study plot is the signature visualization for DiD designs. Pre-treatment coefficients near zero validate the parallel trends assumption, while post-treatment coefficients reveal the dynamic treatment effect:
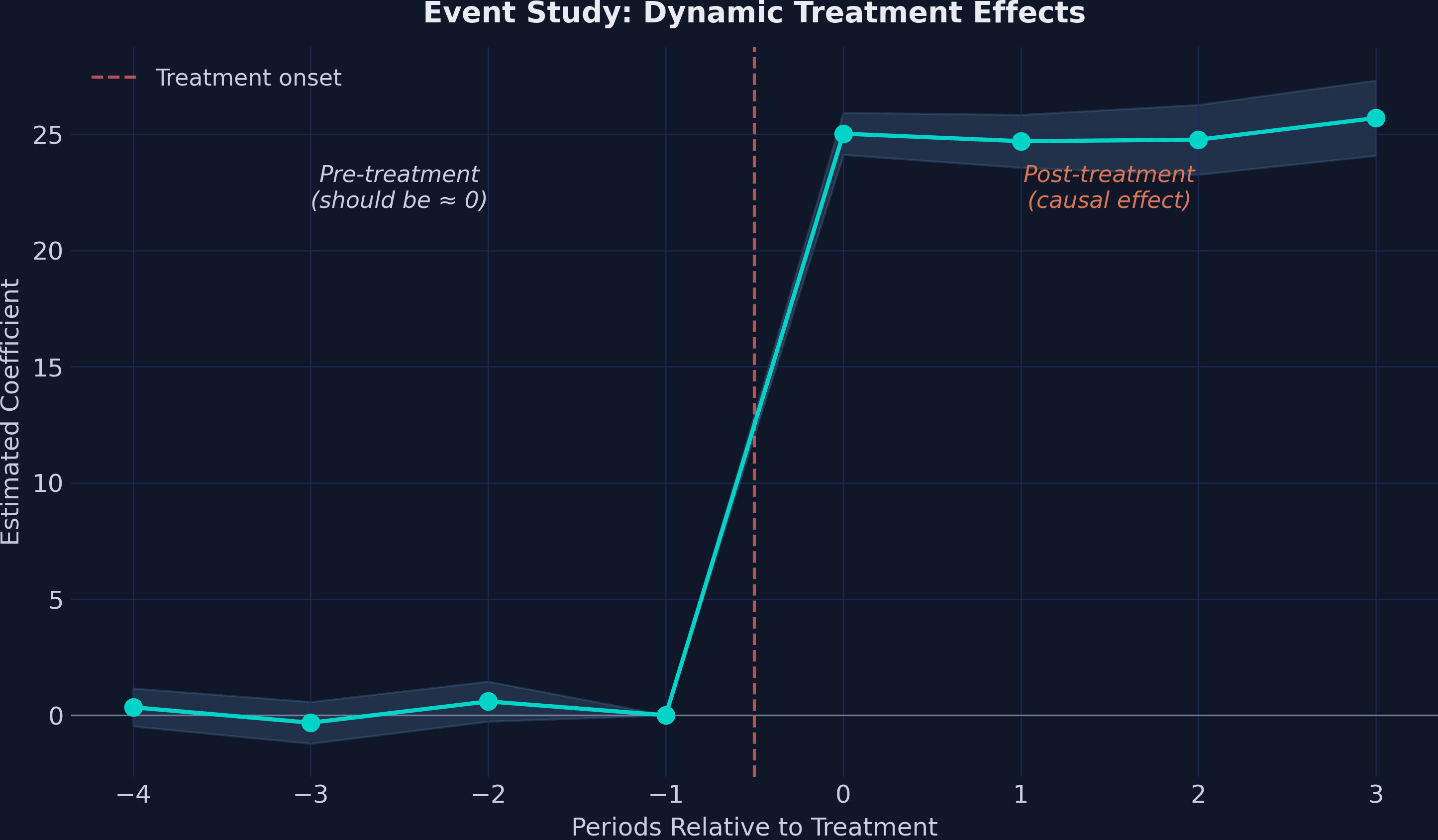
The plot shows a textbook event study pattern:
-
Pre-treatment (t = −4 to −2): Coefficients are small (0.34, −0.32, 0.59) and statistically insignificant (all p > 0.17). The confidence intervals comfortably include zero. This is strong evidence supporting the parallel trends assumption — treated and comparison schools were following similar trajectories before the program.
-
Post-treatment (t = 0 to 3): An immediate, sharp jump to ≈25 points at the moment of treatment, with the effect remaining remarkably stable across all four post-treatment periods (24.71 to 25.70). There is no evidence of fade-out or dynamic adjustment — the program’s effect is both immediate and sustained.
11.5 Event study coefficients table
| Period | Estimate | 95% CI | Significant? |
|---|---|---|---|
| t = −4 | 0.342 | [−0.47, 1.16] | No |
| t = −3 | −0.322 | [−1.22, 0.57] | No |
| t = −2 | 0.593 | [−0.27, 1.45] | No |
| t = −1 | 0.000 | (reference) | — |
| t = 0 | 25.028 | [24.12, 25.93] | Yes (p < 0.001) |
| t = 1 | 24.705 | [23.57, 25.84] | Yes (p < 0.001) |
| t = 2 | 24.768 | [23.27, 26.27] | Yes (p < 0.001) |
| t = 3 | 25.701 | [24.08, 27.32] | Yes (p < 0.001) |
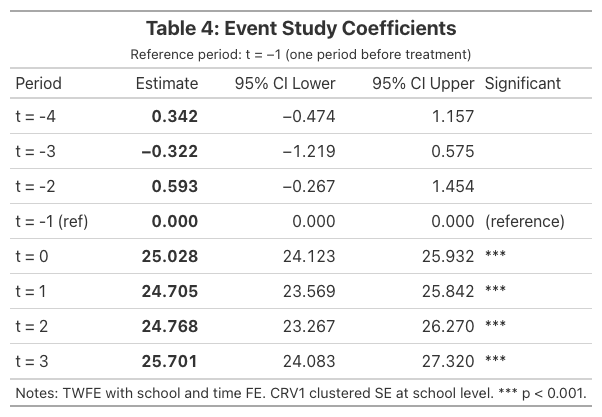
The event study confirms three critical findings: (1) no pre-trends — the design is credible, (2) immediate effect — the program works from day one, and (3) sustained impact — no fade-out over four post-treatment periods.
12. Discussion
12.1 Four key findings
-
The naive before-after comparison overstates the effect by 43%. The raw change in treated schools is 36.20 GPA points, but 10.88 of these points reflect a common upward trend shared by all schools. DiD correctly attributes only 25.32 points to the program.
-
The event study confirms no differential pre-trends. All three pre-treatment coefficients (0.34, −0.32, 0.59) are small, close to zero, and statistically insignificant. The parallel trends assumption is well-supported by the data.
-
The effect is immediate and sustained. Post-treatment coefficients range from 24.71 to 25.70, showing no evidence of delayed onset, gradual ramp-up, or fade-out. The tutoring program’s impact is both immediate and remarkably stable.
-
Inference choice matters less than design. Standard errors ranged from 0.585 (CRV1) to 0.637 (CRV3) across four inference methods, but all produced t-statistics above 39. When the research design is clean and the signal is strong, the choice of variance estimator is practically irrelevant.
12.2 Caveats
This tutorial uses simulated data designed to illustrate DiD mechanics cleanly. In real applications, you should expect:
- R-squared below 0.99: The R² of 0.995 in our TWFE models is unrealistically high. Real education data has far more noise.
- Smaller treatment effects: A 25-point GPA increase is enormous. Real programs typically produce single-digit effects.
- Imperfect parallel trends: Pre-treatment coefficients may not be exactly zero, requiring judgment about how much deviation is acceptable.
- Staggered treatment timing: When different units receive treatment at different times, the standard TWFE estimator can be biased. Modern DiD estimators (Callaway & Sant’Anna, 2021; Gardner, 2022) address this.
13. Summary and Takeaways
-
DiD removes common time trends. The naive approach overstated the effect by 10.88 GPA points — exactly the secular trend captured by the comparison group.
-
Multiple approaches produce one answer. Three specifications (OLS, TWFE, TWFE with covariate) all yield a DiD estimate of 25.32–25.33, demonstrating the robustness of the design.
-
Event studies test parallel trends. Pre-treatment coefficients (0.34, −0.32, 0.59) are close to zero and insignificant, providing strong evidence that treated and comparison schools were on similar trajectories before the program.
-
The effect is immediate and sustained. Post-treatment coefficients (24.71–25.70) show no evidence of delayed onset or fade-out.
-
Inference flexibility is a PyFixest strength. Switching between iid, HC1, CRV1, and CRV3 requires only changing the
vcovargument — no need for separate packages or commands. -
etable() and Great Tables replace manual table construction. The
csw0()operator estimates multiple specifications in one call, andetable()produces publication-ready output. For custom formatting, Great Tables provides full control via.tab_header(),.fmt_number(),.tab_style(), and.save().
14. Exercises
-
Robustness check: Load the event study dataset and collapse it to a 2×2 design by averaging GPA across all pre-treatment periods and all post-treatment periods for each school. Re-estimate the DiD. Does the estimate match the 2×2 result?
-
Inference sensitivity: Estimate the TWFE model with all four SE types (iid, HC1, CRV1, CRV3). At what significance level (α) would the choice of SE type change your conclusion about statistical significance? How few clusters would you need before the choice starts to matter?
-
Staggered DiD: Read about PyFixest’s
did2s()function for the Gardner (2022) two-stage DiD estimator. How would you adapt this tutorial’s code to handle a setting where the 10 treated schools received the program at different times?
References
- Corral, D. & Yang, M. (2024). An introduction to the difference-in-differences design in education policy research. Asia Pacific Education Review.
- Callaway, B. & Sant’Anna, P.H. (2021). Difference-in-differences with multiple time periods. Journal of Econometrics, 225(2), 200–230.
- Goodman-Bacon, A. (2021). Difference-in-differences with variation in treatment timing. Journal of Econometrics, 225(2), 254–277.
- Sun, L. & Abraham, S. (2021). Estimating dynamic treatment effects in event studies with heterogeneous treatment effects. Journal of Econometrics, 225(2), 175–199.
- Borusyak, K., Jaravel, X. & Spiess, J. (2024). Revisiting event-study designs: robust and efficient estimation. Review of Economic Studies, 91(6), 3253–3285.
- Baker, A.C., Larcker, D.F. & Wang, C.C.Y. (2022). How much should we trust staggered difference-in-differences estimates? Journal of Financial Economics, 144(2), 370–395.
- Correia, S. (2016). REGHDFE: Stata module to perform linear or instrumental-variable regression absorbing any number of high-dimensional fixed effects.
- Fischer, A. & Schar, A. (2024). PyFixest: Fast high-dimensional fixed effects estimation in Python.
- Great Tables: Presentation-ready display tables. Posit PBC.
- Gardner, J. (2022). Two-stage differences in differences. Working paper.