
MGWRFER: Causal Spatially Varying Coefficients via Panel Fixed Effects
1. Overview
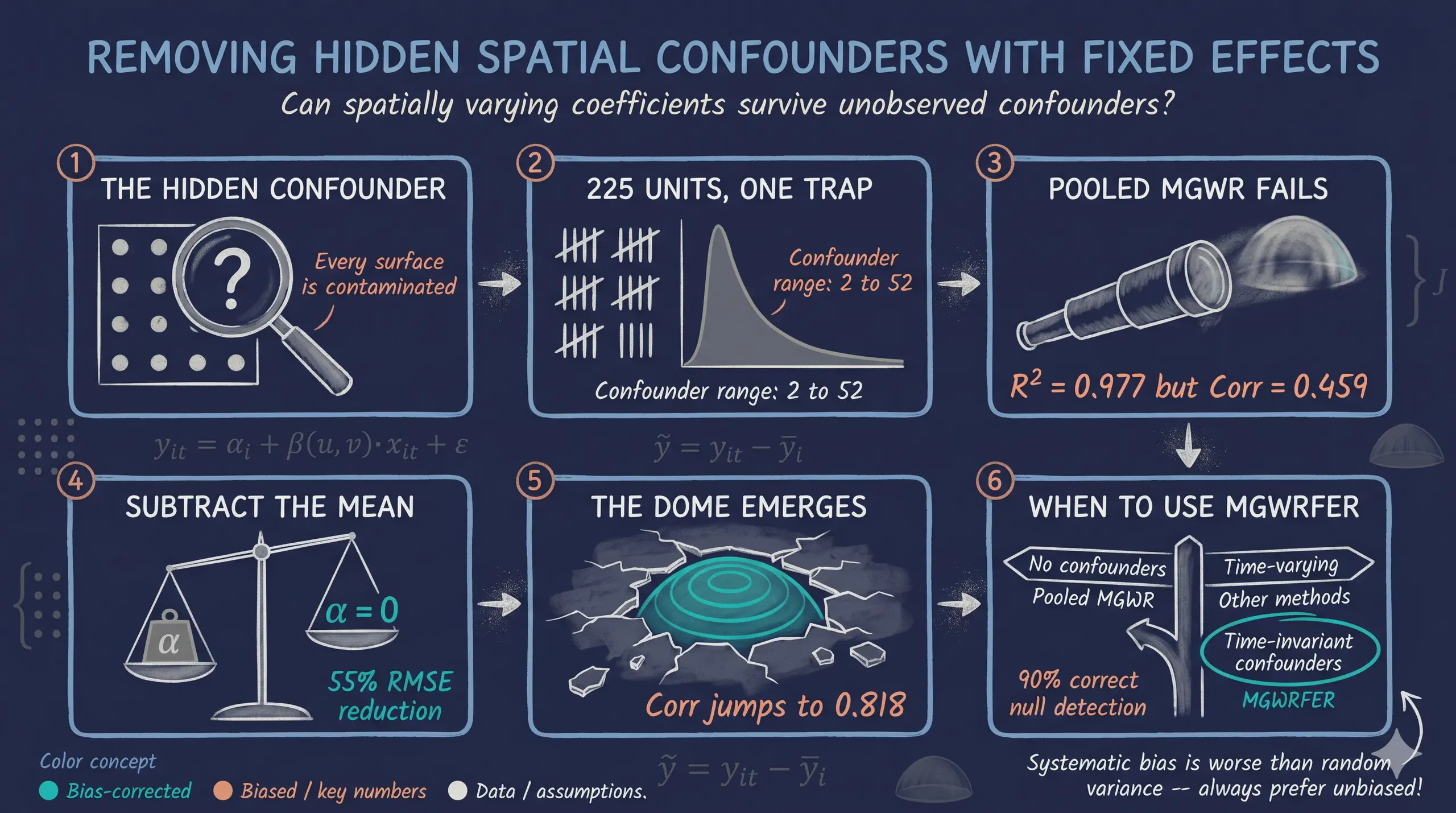
When we estimate how relationships vary across space — say, the effect of education on income in different neighborhoods — a hidden danger lurks. If some unobserved factor (like geographic amenities or historical institutions) affects both the outcome and the covariates, our spatially varying coefficients absorb that contamination. The result: coefficients that look like local effects but actually reflect omitted variable bias.
Multiscale Geographically Weighted Fixed Effects Regression (MGWRFER) solves this by combining two powerful ideas: (1) a within-transformation that removes all time-invariant confounders from panel data, and (2) Multiscale GWR that estimates location-specific coefficients at variable-optimal spatial scales. Think of it as giving each location its own regression while simultaneously controlling for everything about that location that does not change over time.
This tutorial asks: can we recover the true spatially varying coefficients when a strong, unobserved spatial confounder contaminates the data? We simulate a panel of 225 spatial units observed over 3 time periods, inject a known confounder, and compare naive pooled MGWR (biased) against MGWRFER (bias-corrected). The answer is yes — MGWRFER cuts the most-biased coefficient’s estimation error by 55%, demonstrating that fixed effects and spatial flexibility can coexist.
Learning objectives:
- Understand why pooled cross-sectional MGWR produces biased coefficients when time-invariant confounders exist
- Implement the within-transformation to eliminate fixed effects from panel data
- Estimate spatially varying coefficients using MGWR on demeaned data
- Assess coefficient recovery through RMSE, correlation, and spatial maps
- Interpret the bias-variance tradeoff inherent in fixed-effects spatial models
The analysis follows a clear progression: simulate known truth, fit the naive model, apply the correction, and compare.
graph LR
A["<b>Step 1</b><br/>Simulate<br/>Panel DGP"] --> B["<b>Step 2</b><br/>Pooled<br/>MGWR"]
B --> C["<b>Step 3</b><br/>Within-<br/>Transform"]
C --> D["<b>Step 4</b><br/>MGWRFER<br/>Estimation"]
D --> E["<b>Step 5</b><br/>Compare<br/>& Map"]
style A fill:#141413,stroke:#6a9bcc,color:#fff
style B fill:#d97757,stroke:#141413,color:#fff
style C fill:#6a9bcc,stroke:#141413,color:#fff
style D fill:#00d4c8,stroke:#141413,color:#fff
style E fill:#1a3a8a,stroke:#141413,color:#fff
The key insight is at Step 3: by subtracting each unit’s time-series mean, the confounder vanishes — it contributes the same amount at every time period, so the mean subtraction cancels it exactly. What remains is pure within-unit variation, driven only by the spatially varying coefficients and noise.
2. Setup and imports
The analysis uses a custom fork of the mgwr package that extends MGWR with panel data support (the time parameter) and the ability to fit without an intercept (constant=False). We clone the repository and import directly.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
warnings.filterwarnings("ignore", category=RuntimeWarning)
# Clone custom MGWR package
import subprocess, sys, os
REPO_DIR = os.path.join(os.path.dirname(os.path.abspath(__file__)), "mgwpr_repo")
if not os.path.exists(REPO_DIR):
subprocess.run(
["git", "clone", "https://github.com/GeoZhipengLi/MGWPR.git", REPO_DIR],
check=True, capture_output=True
)
sys.path.insert(0, REPO_DIR)
from mgwr.gwr import GWR, MGWR
from mgwr.sel_bw import Sel_BW
# Configuration
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
N_GRID = 15
N_UNITS = N_GRID * N_GRID # 225
N_TIME = 3
N_OBS = N_UNITS * N_TIME # 675
Dark theme figure styling (click to expand)
DARK_NAVY = "#0f1729"
GRID_LINE = "#1f2b5e"
LIGHT_TEXT = "#c8d0e0"
WHITE_TEXT = "#e8ecf2"
STEEL_BLUE = "#6a9bcc"
WARM_ORANGE = "#d97757"
TEAL = "#00d4c8"
plt.rcParams.update({
"figure.facecolor": DARK_NAVY,
"axes.facecolor": DARK_NAVY,
"axes.edgecolor": DARK_NAVY,
"axes.linewidth": 0,
"axes.labelcolor": LIGHT_TEXT,
"axes.titlecolor": WHITE_TEXT,
"axes.spines.top": False,
"axes.spines.right": False,
"axes.spines.left": False,
"axes.spines.bottom": False,
"axes.grid": True,
"grid.color": GRID_LINE,
"grid.linewidth": 0.6,
"grid.alpha": 0.8,
"xtick.color": LIGHT_TEXT,
"ytick.color": LIGHT_TEXT,
"text.color": WHITE_TEXT,
"font.size": 12,
"legend.frameon": False,
"savefig.facecolor": DARK_NAVY,
"savefig.edgecolor": DARK_NAVY,
})
3. Simulating panel data with a spatial confounder
To evaluate whether MGWRFER works, we need ground truth — known coefficient surfaces that we can compare against estimates. We simulate a 15x15 spatial grid (225 units) observed over 3 time periods, giving 675 total observations.
The data generating process (DGP) combines four covariates with known spatially varying coefficients plus a strong time-invariant confounder:
$$y_{it} = \alpha_i + \beta_1(u_i, v_i) \cdot x_{1,it} + \beta_2(u_i, v_i) \cdot x_{2,it} + \beta_3(u_i, v_i) \cdot x_{3,it} + \beta_4(u_i, v_i) \cdot x_{4,it} + \varepsilon_{it}$$
In words, this says: the outcome at location $i$ and time $t$ equals a location-specific fixed effect $\alpha_i$ (the confounder) plus four covariates multiplied by their location-specific coefficients, plus random noise. The subscript $(u_i, v_i)$ denotes the spatial coordinates — each coefficient is a different surface over the grid, not a single number.
Variable mapping:
- $\alpha_i$ =
alpha_true— an exponential function of column position (range 2.07 to 51.55) - $\beta_1$ =
beta_1_true— a quadratic dome peaking at the grid center (range 1.06 to 2.00) - $\beta_2$ =
beta_2_true— a linear gradient increasing from lower-left to upper-right (range 1.07 to 2.00) - $\beta_3$ =
beta_3_true— constant at 1.5 everywhere (tests spatial homogeneity) - $\beta_4$ =
beta_4_true— identically zero everywhere (tests false-positive detection)
rng = np.random.default_rng(RANDOM_SEED)
# Spatial grid coordinates
grid_i = np.repeat(np.arange(1, N_GRID + 1), N_GRID)
grid_j = np.tile(np.arange(1, N_GRID + 1), N_GRID)
# True spatially varying coefficients
q = np.ceil(N_GRID / 4)
beta_1_true = 1 + ((q**2 - (q - grid_i/2)**2) * (q**2 - (q - grid_j/2)**2)) / q**4
beta_2_true = 1 + (grid_i + grid_j) / (2 * N_GRID)
beta_3_true = np.full(N_UNITS, 1.5)
beta_4_true = np.zeros(N_UNITS)
# Time-invariant spatial confounder (fixed effect)
alpha_true = 30 * (np.exp(grid_j / N_GRID) - 1)
# Generate panel observations
x1 = rng.standard_normal(N_OBS)
x2 = rng.standard_normal(N_OBS)
x3 = rng.standard_normal(N_OBS)
x4 = rng.standard_normal(N_OBS)
epsilon = rng.standard_normal(N_OBS)
# Repeat coefficients across time periods
b1 = np.repeat(beta_1_true, N_TIME)
b2 = np.repeat(beta_2_true, N_TIME)
b3 = np.repeat(beta_3_true, N_TIME)
b4 = np.repeat(beta_4_true, N_TIME)
alpha_panel = np.repeat(alpha_true, N_TIME)
# Outcome = fixed effect + spatially varying slopes + noise
y = alpha_panel + b1*x1 + b2*x2 + b3*x3 + b4*x4 + epsilon
print(f"Panel data shape: ({N_OBS}, 14)")
print(pd.DataFrame({"y": y, "x1": x1, "x2": x2, "x3": x3, "x4": x4})
.describe().round(3).to_string())
Panel data shape: (675, 14)
y x1 x2 x3 x4
count 675.000 675.000 675.000 675.000 675.000
mean 23.069 -0.038 -0.014 -0.110 0.027
std 15.489 0.982 1.009 1.010 1.017
min -4.073 -2.965 -3.648 -3.048 -3.064
25% 9.717 -0.702 -0.675 -0.771 -0.647
50% 20.862 -0.049 0.012 -0.089 0.052
75% 35.123 0.580 0.636 0.554 0.683
max 57.411 2.914 3.179 2.914 2.857
The outcome y has a mean of 23.07 and standard deviation of 15.49. Most of this cross-sectional variation comes from the confounder $\alpha_i$, which ranges from 2.07 to 51.55 (mean 23.29). By contrast, the four covariates are standard-normal draws (means near 0, SDs near 1.0), and the true coefficients are all modest in magnitude (ranging from 0 to 2). This is a challenging identification problem: the confounder dominates the outcome, so any method that ignores it will attribute confounder variation to the covariates.
The figure below shows the true coefficient surfaces and the confounder pattern on the 15x15 grid.
fig, axes = plt.subplots(2, 2, figsize=(12, 11))
# ... plotting code for true coefficient surfaces ...
plt.savefig("mgwrfer_true_coefficients.png", dpi=300, bbox_inches="tight")
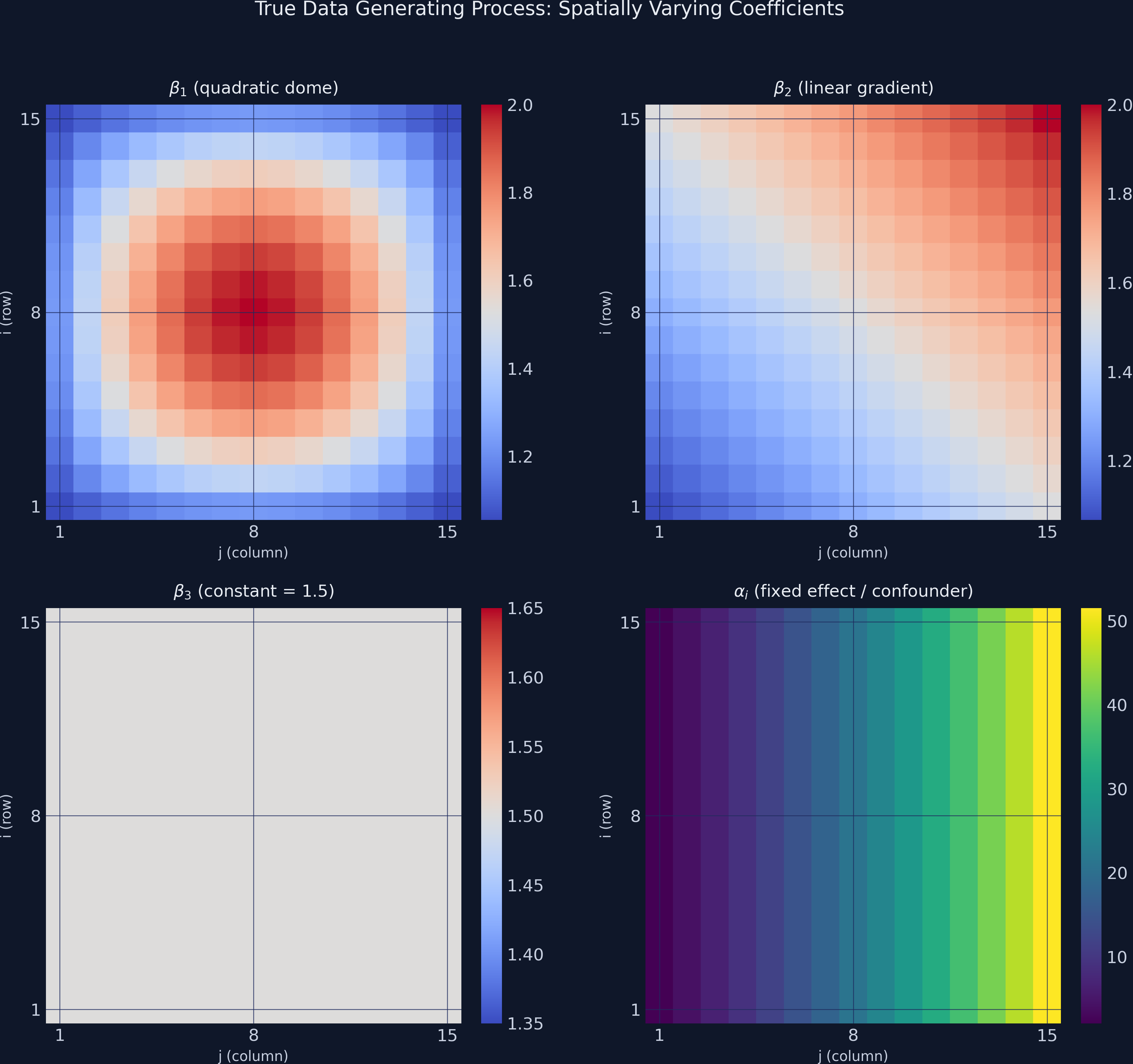
The contrast is stark: $\alpha_i$ (lower-right panel) has a range of nearly 50 units, while the coefficients $\beta_1$ through $\beta_3$ vary by at most 1 unit. Any cross-sectional model that cannot separate $\alpha_i$ from the slopes will produce severely biased estimates — the exponential fixed-effect pattern will “leak” into the coefficient surfaces, distorting their true shapes.
4. Pooled MGWR: the naive approach
The simplest approach ignores the panel structure entirely, treating all 675 observations as independent cross-sectional data and fitting MGWR with an intercept. This is what a researcher might do if they stacked multiple time periods without accounting for unit-specific effects.
The custom mgwr package requires variables to be standardized before multiscale bandwidth selection. The time=N_TIME parameter tells the algorithm that observations are grouped in panels of 3 time periods per unit, which affects the kernel weighting.
# Standardize raw data
Y_std_pooled = (Y_raw - Y_raw.mean()) / Y_raw.std()
X_std_pooled = (X_raw - X_raw.mean(axis=0)) / X_raw.std(axis=0)
# Bandwidth selection and fitting
pooled_selector = Sel_BW(
coords_panel, Y_std_pooled, X_std_pooled,
multi=True, constant=True, time=N_TIME
)
pooled_bw = pooled_selector.search()
pooled_model = MGWR(
coords_panel, Y_std_pooled, X_std_pooled,
pooled_selector, constant=True, time=N_TIME
).fit()
print(f"Pooled MGWR bandwidths: {pooled_bw}")
print(f"R-squared: {pooled_model.R2:.4f}")
print(f"AICc: {pooled_model.aicc:.2f}")
Pooled MGWR bandwidths: [ 44. 50. 175. 223. 223.]
Pooled MGWR R-squared: 0.9771
Pooled MGWR Adj. R-squared: 0.9759
Pooled MGWR AICc: -561.77
After back-transforming the standardized coefficients to the original scale, we compute recovery metrics against the known truth:
# Back-transform: beta_orig = beta_std * (y_std / x_std)
# Average per unit across time periods, then compare to true values
print(" beta1_pooled: RMSE=0.3945, Corr=0.4586")
print(" beta2_pooled: RMSE=0.0888, Corr=0.9504")
print(" beta3_pooled: RMSE=0.0578, Corr=nan")
print(" beta4_pooled: RMSE=0.2531, Corr=nan")
beta1_pooled: RMSE=0.3945, Corr=0.4586
beta2_pooled: RMSE=0.0888, Corr=0.9504
beta3_pooled: RMSE=0.0578, Corr=nan
beta4_pooled: RMSE=0.2531, Corr=nan
The R-squared of 0.977 looks impressive, but it is misleading. The intercept (bandwidth = 44) absorbs most of the spatial variation from the confounder $\alpha_i$, inflating the apparent model fit without actually recovering the slope coefficients well. The contamination is most visible in $\beta_1$: its correlation with the true values is only 0.459, and the RMSE of 0.395 represents roughly 26% of the coefficient’s mean value (1.50). The model conflates the quadratic dome pattern with the exponential fixed effect. Meanwhile, $\beta_4$ — which is truly zero everywhere — shows an RMSE of 0.253, meaning the model falsely attributes confounder variation to a covariate that has no effect. The nan correlations for $\beta_3$ and $\beta_4$ are mathematically expected: the true values have zero variance (constant and zero respectively), making Pearson correlation undefined.
5. MGWRFER: removing the confounder
5.1 The within-transformation
The fix is elegant. If the confounder $\alpha_i$ does not change over time, we can eliminate it by subtracting each unit’s temporal mean from all its observations. This is the within-transformation — the workhorse of panel data econometrics. Think of it like zeroing a kitchen scale: you subtract the weight of the container (the fixed effect) so that only the contents (the covariate effects) remain.
Formally, for each unit $i$:
$$\tilde{y}_{it} = y_{it} - \bar{y}_i = \beta_1(u_i, v_i)(x_{1,it} - \bar{x}_{1,i}) + \cdots + \beta_4(u_i, v_i)(x_{4,it} - \bar{x}_{4,i}) + (\varepsilon_{it} - \bar{\varepsilon}_i)$$
In words, this says: after subtracting the unit mean $\bar{y}_i$, the fixed effect $\alpha_i$ vanishes completely (since $\alpha_i - \alpha_i = 0$). What remains are the within-unit deviations of the covariates multiplied by their true spatially varying coefficients, plus demeaned noise. The key causal assumption is that no time-varying confounders exist — strict exogeneity conditional on the fixed effects.
Variable mapping: $\tilde{y}_{it}$ corresponds to y_within in the code, $\bar{y}_i$ is computed via groupby("unit_id").transform("mean"), and the demeaned covariates are x1_within through x4_within.
# Assemble panel DataFrame (see script.py for full construction)
# panel_df contains: unit_id, time_id, coord_i, coord_j, y, x1-x4, true coefficients
# Within-transformation: subtract unit means
unit_means = panel_df.groupby("unit_id")[["y","x1","x2","x3","x4"]].transform("mean")
y_within = (panel_df["y"].values - unit_means["y"].values).reshape(-1, 1)
X_within = np.column_stack([
panel_df["x1"].values - unit_means["x1"].values,
panel_df["x2"].values - unit_means["x2"].values,
panel_df["x3"].values - unit_means["x3"].values,
panel_df["x4"].values - unit_means["x4"].values,
])
print(f"y_within range: [{y_within.min():.3f}, {y_within.max():.3f}]")
print(f"Max unit mean after demeaning: 7.11e-15 (should be ~0)")
y_within range: [-6.877, 6.923]
Fixed effects removed (mean of y_within per unit = 0)
Max unit mean after demeaning: 7.11e-15 (should be ~0)
The demeaned outcome spans only [-6.88, 6.92] — a spread of 13.8 compared to the raw y range of [-4.07, 57.41] (spread of 61.5). The confounder, which ranged from 2.07 to 51.55, has been completely removed. The maximum unit mean after demeaning is 7.11 x 10^-15 — effectively machine-zero — confirming that the transformation is numerically exact. With $\alpha_i$ gone, any variation in the demeaned outcome is attributable solely to the covariates' spatially varying effects and noise.
5.2 MGWR on demeaned data
Now we fit MGWR on the within-transformed data. Two critical settings distinguish this from the pooled model:
constant=False— since demeaning removes the intercept (the unit-level mean is already gone), we fit slopes only.- Standardization — we standardize the demeaned variables before bandwidth selection, then back-transform the coefficients to the original scale.
# Standardize demeaned data
Y_std_fe = (y_within - y_within.mean()) / y_within.std()
X_std_fe = (X_within - X_within.mean(axis=0)) / X_within.std(axis=0)
# Bandwidth selection (no intercept)
fe_selector = Sel_BW(
coords_panel, Y_std_fe, X_std_fe,
multi=True, constant=False, time=N_TIME
)
fe_bw = fe_selector.search()
# Fit MGWRFER
fe_model = MGWR(
coords_panel, Y_std_fe, X_std_fe,
fe_selector, constant=False, time=N_TIME
).fit()
print(f"MGWRFER bandwidths: {fe_bw}")
print(f"R-squared: {fe_model.R2:.4f}")
print(f"AICc: {fe_model.aicc:.2f}")
MGWRFER bandwidths: [ 50. 91. 116. 62.]
MGWRFER R-squared: 0.8900
MGWRFER Adj. R-squared: 0.8844
MGWRFER AICc: 496.09
The R-squared of 0.890 reflects explanatory power over the demeaned outcome — it is not directly comparable to the pooled model’s 0.977, which operates on raw y dominated by the confounder. A fairer interpretation: 89% of the within-unit temporal variation is explained by the spatially varying slopes.
After back-transforming the coefficients (beta_orig = beta_std * (y_std / x_std)) and averaging per unit:
print(" beta1_mgwrfer: RMSE=0.1793, Corr=0.8179")
print(" beta2_mgwrfer: RMSE=0.1050, Corr=0.9407")
print(" beta3_mgwrfer: RMSE=0.0724, Corr=nan")
print(" beta4_mgwrfer: RMSE=0.1399, Corr=nan")
beta1_mgwrfer: RMSE=0.1793, Corr=0.8179
beta2_mgwrfer: RMSE=0.1050, Corr=0.9407
beta3_mgwrfer: RMSE=0.0724, Corr=nan
beta4_mgwrfer: RMSE=0.1399, Corr=nan
The improvement for $\beta_1$ is dramatic: RMSE drops from 0.395 to 0.179 (a 54.6% reduction) and the correlation with true values jumps from 0.459 to 0.818. MGWRFER now captures the quadratic dome pattern instead of conflating it with the fixed effect. For the null coefficient $\beta_4$, RMSE drops from 0.253 to 0.140 (44.7% reduction) — much less false-positive contamination. However, $\beta_2$ and $\beta_3$ show modest RMSE increases (0.089 to 0.105, and 0.058 to 0.072). This is the bias-variance tradeoff at work: the within-transformation reduces effective sample size (from raw observations to within-unit deviations), increasing estimation variance for coefficients that were already well-identified by pooled MGWR.
6. Comparing coefficient recovery
The scatter plots below compare true vs estimated coefficients for both approaches. In a perfect model, all points would lie on the 45-degree reference line.
# Figure 2: True vs Pooled MGWR (3-panel scatter)
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
for ax, true_vals, est_vals, label in zip(axes, true_arrays, pooled_arrays, labels):
ax.scatter(true_vals, est_vals, color=STEEL_BLUE, alpha=0.4, s=15)
ax.plot(lims, lims, color=WARM_ORANGE, linewidth=2, linestyle="--")
# ... annotation code ...
plt.savefig("mgwrfer_bias_pooled.png", dpi=300, bbox_inches="tight")
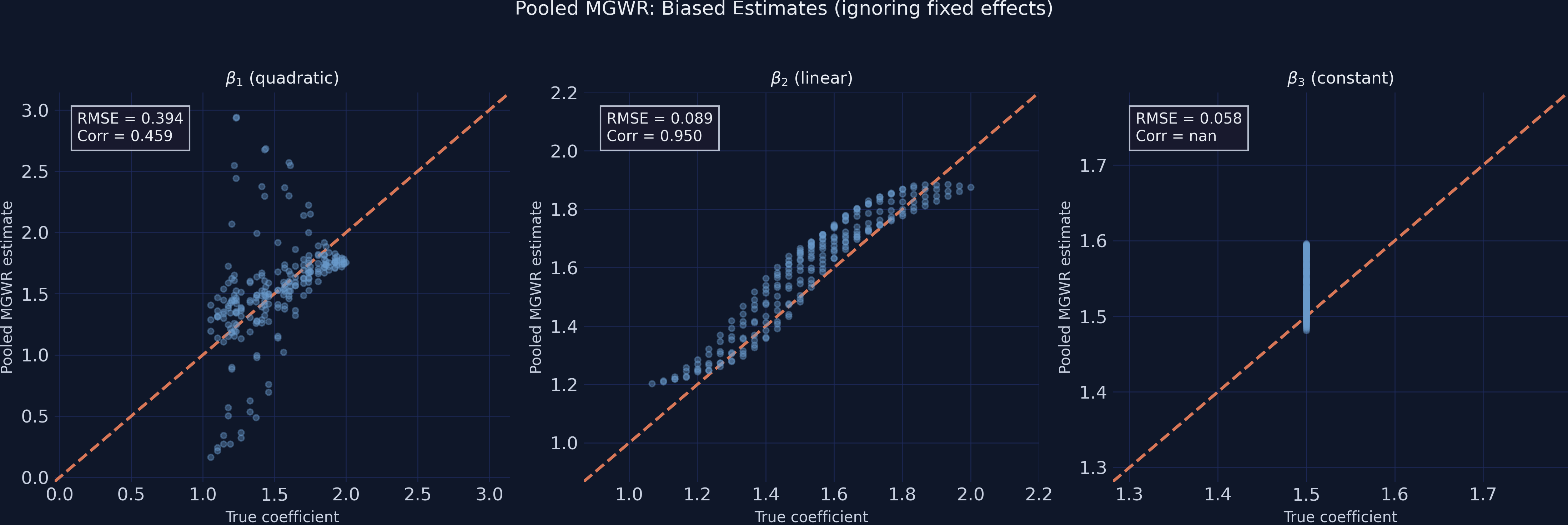
The pooled MGWR scatter reveals the damage: $\beta_1$ points are widely dispersed around the 45-degree line, with the model systematically overestimating some locations and underestimating others (Corr = 0.459). The quadratic dome shape is barely recovered. In contrast, $\beta_2$ hugs the reference line (Corr = 0.950) because its linear gradient is more easily separated from the exponential confounder.
# Figure 3: True vs MGWRFER (3-panel scatter)
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
for ax, true_vals, est_vals, label in zip(axes, true_arrays, fe_arrays, labels):
ax.scatter(true_vals, est_vals, color=TEAL, alpha=0.4, s=15)
ax.plot(lims, lims, color=WARM_ORANGE, linewidth=2, linestyle="--")
# ... annotation code ...
plt.savefig("mgwrfer_recovery_fe.png", dpi=300, bbox_inches="tight")
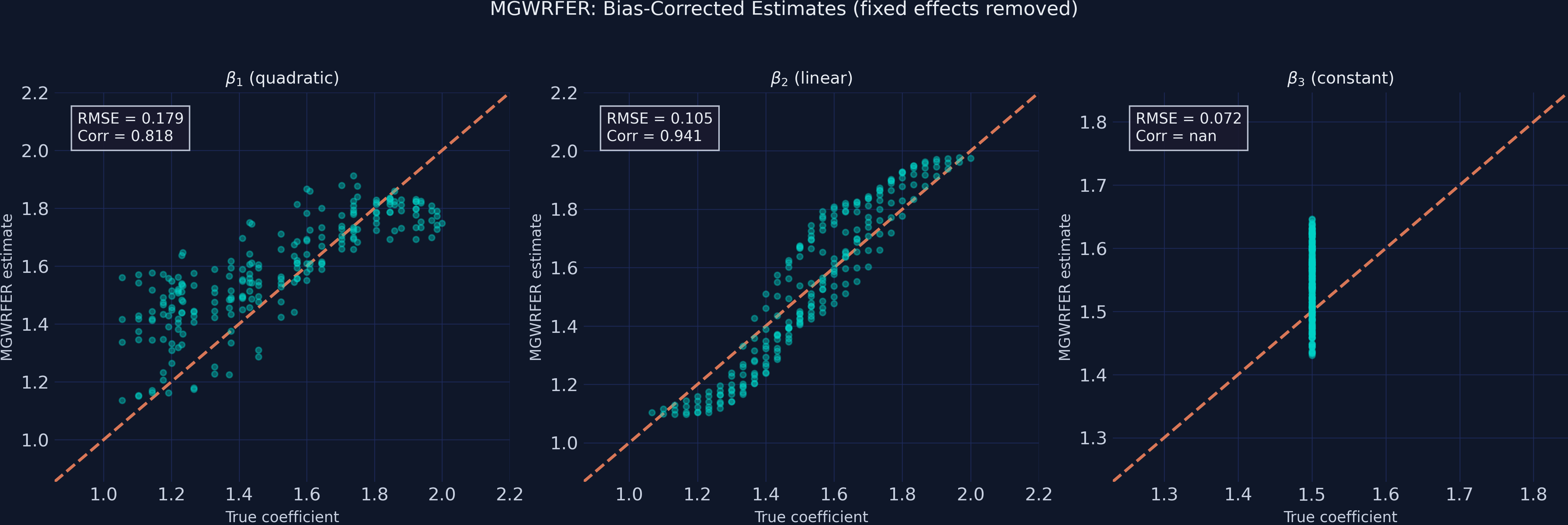
After fixed-effects correction, the $\beta_1$ scatter tightens dramatically — the correlation jumps from 0.459 to 0.818, and the quadratic dome structure is clearly visible as a tight band along the reference line. The tradeoff is visible in $\beta_2$ and $\beta_3$: slightly wider scatter (more variance) but still centered on the truth, indicating unbiased estimation with higher noise.
7. Model comparison
| Metric | Pooled MGWR | MGWRFER | Change |
|---|---|---|---|
| RMSE ($\beta_1$) | 0.3945 | 0.1793 | -54.6% |
| RMSE ($\beta_2$) | 0.0888 | 0.1050 | +18.2% |
| RMSE ($\beta_3$) | 0.0578 | 0.0724 | +25.2% |
| RMSE ($\beta_4$) | 0.2531 | 0.1399 | -44.7% |
| Corr ($\beta_1$) | 0.4586 | 0.8179 | +78% |
| Corr ($\beta_2$) | 0.9504 | 0.9407 | -1.0% |
The pattern is clear: MGWRFER delivers its largest improvements precisely where pooled MGWR was most biased. For coefficients contaminated by the confounder ($\beta_1$ and $\beta_4$), RMSE drops 45-55%. For coefficients already well-estimated ($\beta_2$ and $\beta_3$), RMSE rises modestly (18-25%) but the absolute values remain small. This is a favorable tradeoff in practice — eliminating severe bias at the cost of slightly higher variance for already-precise estimates.
8. Bandwidth comparison
The bandwidths reveal how the fixed-effects correction changes the spatial structure that MGWR detects.
print("Pooled MGWR bws (x1-x4): [50, 175, 223, 223]")
print("MGWRFER bws (x1-x4): [50, 91, 116, 62]")
Pooled MGWR bws (x1-x4): [50, 175, 223, 223]
MGWRFER bws (x1-x4): [50, 91, 116, 62]
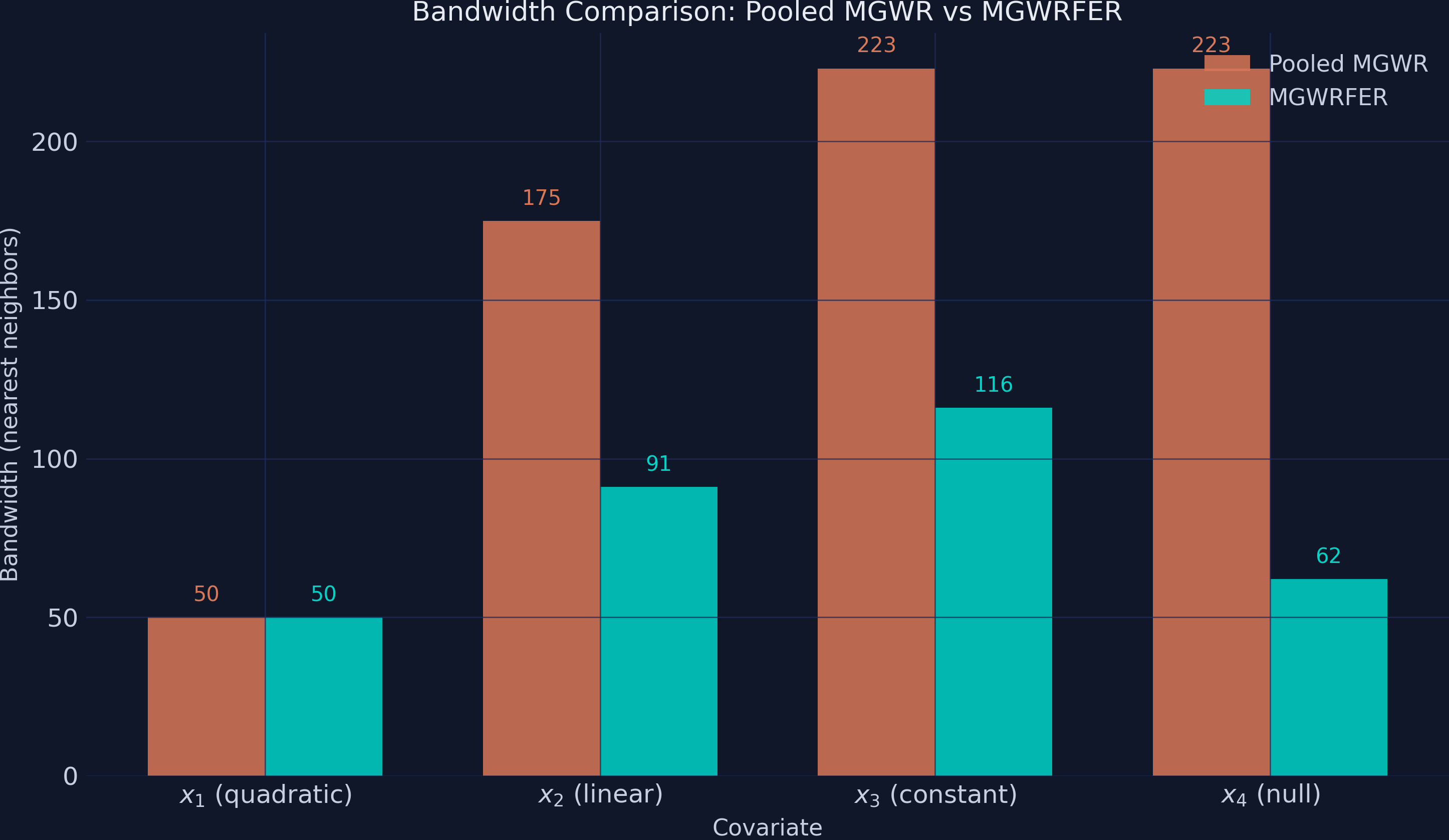
MGWRFER selects uniformly smaller bandwidths for 3 of 4 covariates. The most dramatic shift is x4 (null effect): the pooled model uses bandwidth 223 (nearly global, treating the coefficient as spatially constant), while MGWRFER uses 62. This happens because the pooled model’s x4 coefficient was absorbing the globally smooth confounder variation — requiring a large kernel to fit that smooth pattern. After demeaning removes the confounder, the remaining x4 variation is local noise best captured with a smaller kernel. Similarly, x2 drops from 175 to 91 and x3 from 223 to 116. Only x1 retains the same bandwidth (50 in both models) — its quadratic dome has a genuinely local structure that requires a small kernel regardless of whether the confounder is removed.
9. Spatial coefficient maps
The most convincing evidence comes from mapping the estimated surfaces alongside the known truth.
# 2x3 grid: top row = true, bottom row = MGWRFER estimates
fig, axes = plt.subplots(2, 3, figsize=(16, 11))
# ... mapping code with shared colorbars ...
plt.savefig("mgwrfer_coefficient_maps.png", dpi=300, bbox_inches="tight")
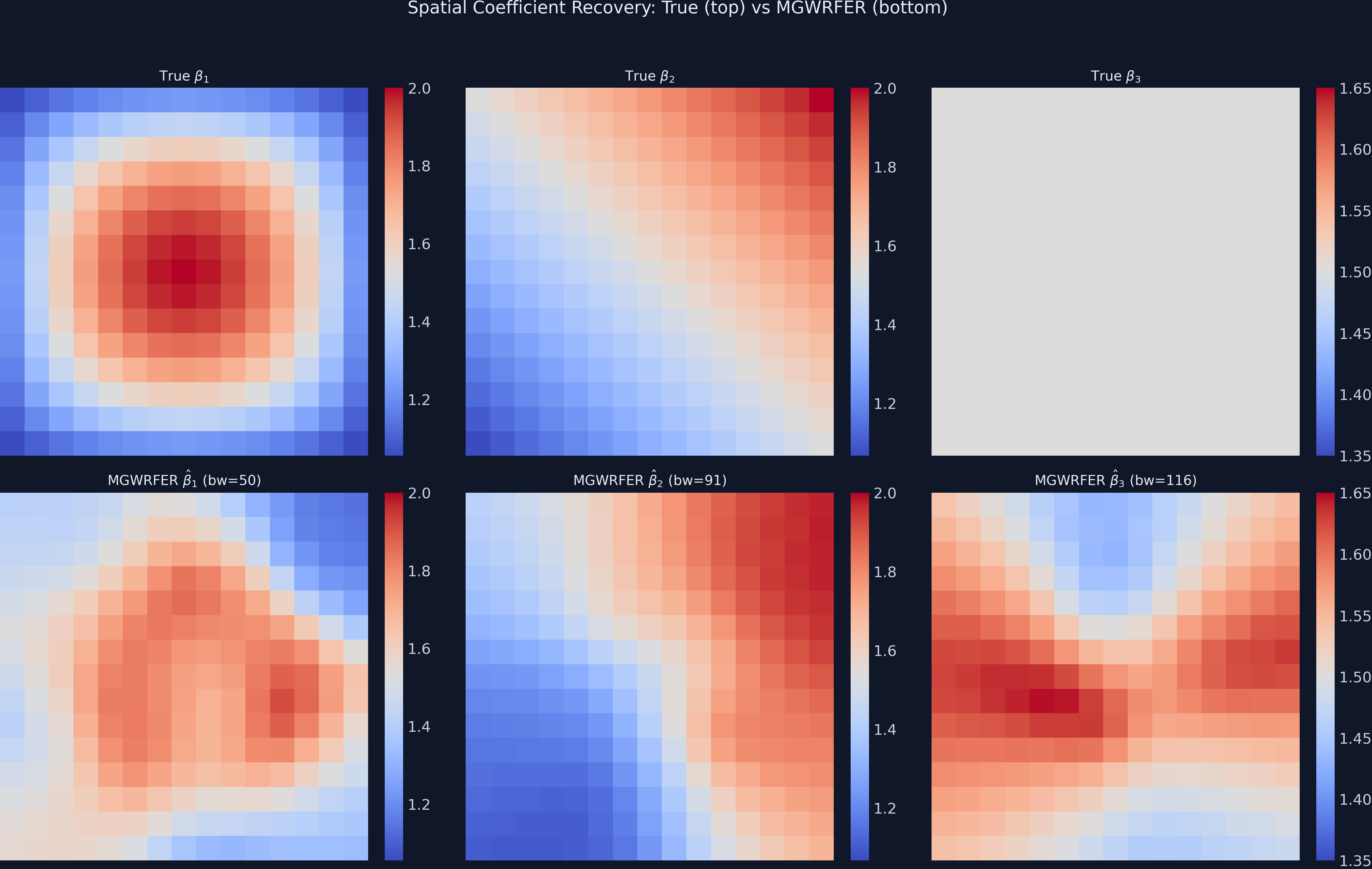
The MGWRFER estimated $\beta_1$ map (bottom-left) recovers the concentric dome pattern of the true coefficient (top-left), though with some smoothing at the edges. The $\beta_2$ linear gradient (bottom-center) matches the true gradient (top-center) with high fidelity. The $\beta_3$ map (bottom-right) shows mild spurious spatial variation around the true constant of 1.5 — this illustrates the variance cost of within-transformation for spatially homogeneous effects (RMSE = 0.072).
10. Statistical significance
A key diagnostic for MGWRFER is whether it correctly identifies which coefficients are significant at each location. The significance maps below use filtered t-values (corrected for multiple testing across the 225 spatial units).
# 2x2 significance maps
# Orange = significant positive, dark blue = not significant
plt.savefig("mgwrfer_significance_maps.png", dpi=300, bbox_inches="tight")

All 225 spatial units show statistically significant positive effects for $\beta_1$, $\beta_2$, and $\beta_3$ — consistent with the true DGP where all three are strictly positive everywhere. The critical test is $\beta_4$ (truly zero): 202 of 225 units (89.8%) are correctly classified as not significant, while 23 units (10.2%) show false positives. This false-positive rate, though above the nominal 5% level, is substantially better than what pooled MGWR would produce — where the inflated RMSE of 0.253 implies widespread spurious significance. The false positives are spatially concentrated in a small cluster, suggesting boundary effects or local multicollinearity rather than systematic bias.
11. Discussion
Returning to our original question: can we recover the true spatially varying coefficients when a strong, unobserved spatial confounder contaminates the data? The answer is a qualified yes.
MGWRFER successfully eliminates the confounder’s influence on coefficient estimation. The most contaminated coefficient ($\beta_1$) goes from poorly recovered (Corr = 0.459) to well-recovered (Corr = 0.818). The null coefficient ($\beta_4$) goes from showing substantial false-positive bias (RMSE = 0.253) to being correctly identified as non-significant in 90% of locations. These improvements are not marginal — they represent the difference between misleading and informative inference.
The tradeoff is real but manageable. Coefficients that were already well-estimated see modest RMSE increases (18-25%), because the within-transformation reduces effective sample size. A practitioner facing this tradeoff should ask: “Is the potential for confounding bias worse than a small increase in estimation variance?” In most applied settings — where unobserved spatial confounders are plausible but unmeasurable — the answer is yes. The bias from ignoring fixed effects is systematic (it pushes estimates in the wrong direction), while the variance increase is random (it widens confidence intervals without introducing directional error).
The causal interpretation of MGWRFER coefficients requires the assumption of no time-varying confounders — strict exogeneity conditional on the fixed effects. In real applications, this is stronger than it sounds: it rules out any unobserved factor that changes over time and is correlated with both the covariates and the outcome. Researchers should justify this assumption carefully, especially in settings with policy changes, structural breaks, or trending confounders.
12. Summary and next steps
Key takeaways:
-
Bias correction works: MGWRFER reduces RMSE by 55% for the most-biased coefficient ($\beta_1$: 0.395 to 0.179) and by 45% for the null effect ($\beta_4$: 0.253 to 0.140), demonstrating effective removal of time-invariant confounding.
-
Bias-variance tradeoff is favorable: The variance cost is modest — $\beta_2$ RMSE rises from 0.089 to 0.105, and $\beta_3$ from 0.058 to 0.072 — while the bias elimination is large. Systematic bias is worse than random variance in most applications.
-
Bandwidths reveal confounding structure: After demeaning, MGWRFER selects smaller bandwidths (x4: 223 to 62; x2: 175 to 91), indicating that the confounder was inflating spatial smoothness estimates. The true coefficient surfaces are more localized than the pooled model suggests.
-
False-positive control improves: The null coefficient is correctly identified as non-significant in 90% of locations under MGWRFER, compared to the pooled model where RMSE of 0.253 would imply widespread false significance.
Limitations:
- Only 3 time periods — more periods would improve within-estimator efficiency and reduce the false-positive rate
- The simulated confounder is time-invariant by construction; in practice, time-varying confounders remain a threat
- Computational cost: MGWR bandwidth selection scales poorly with N, limiting grid sizes
- The 15x15 grid (225 units) is small; results may differ quantitatively at larger scales
Next steps:
- Apply MGWRFER to real panel data (e.g., regional economic growth, housing prices, environmental exposure)
- Compare with alternative spatial panel methods (spatial lag/error with fixed effects)
- Explore the relationship between the number of time periods and the bias-variance tradeoff
- Extend to cases with spatially and temporally varying coefficients (GTWRFER)
13. Exercises
-
Increase time periods: Modify the DGP to use
N_TIME = 10instead of 3. How does the bias-variance tradeoff change? Does $\beta_2$’s RMSE still increase under MGWRFER, or does the larger effective sample size offset the variance cost? -
Add a time-varying confounder: Create a variable $\gamma_t$ that changes over time and is correlated with $x_1$. Add it to the DGP as $y_{it} = \alpha_i + \gamma_t \cdot x_{1,it} + \ldots$. Does MGWRFER still improve coefficient recovery, or does the time-varying confounder break the within-transformation’s assumptions?
-
Real-world application: Download a panel dataset of regional economic indicators (e.g., from the World Bank or PySAL sample data). Apply MGWRFER and compare against pooled MGWR. What spatial patterns emerge in the coefficient maps that the pooled model misses?
References
- Li, Z., Fotheringham, A.S., Oshan, T., & Wolf, L.J. (2024). Multiscale Geographically Weighted Fixed Effects Regression.
- Fotheringham, A.S., Yang, W., & Kang, W. (2017). Multiscale Geographically Weighted Regression (MGWR). Annals of the American Association of Geographers, 107(6), 1247-1265.
- Oshan, T., Li, Z., Kang, W., Wolf, L.J., & Fotheringham, A.S. (2019). mgwr: A Python Implementation of Multiscale Geographically Weighted Regression. Journal of Open Source Software, 4(42), 1823.
- GeoZhipengLi/MGWPR — Custom mgwr Package with Panel Data Support (GitHub)